R中的双圈:使用.name_repair指定修复?
我在R中有这个数据集:
set.seed(123)
myFun <- function(n = 5000) {
a <- do.call(paste0, replicate(5, sample(LETTERS, n, TRUE), FALSE))
paste0(a, sprintf("%04d", sample(9999, n, TRUE)), sample(LETTERS, n, TRUE))
}
col1 = myFun(100)
col2 = myFun(100)
col3 = myFun(100)
col4 = myFun(100)
group <- c("A","B","C","D")
group = sample(group, 100, replace=TRUE)
example = data.frame(col1, col2, col3, col4, group)
col1 col2 col3 col4 group
1 SKZDZ9876D BTAMF8110T LIBFV6882H ZFIPL4295E A
2 NXJRX7189Y AIZGY5809C HSMIH4556D YJGJP8022H C
3 XPTZB2035P EEKXK0873A PCPNW1021S NMROS4134O A
4 LJMCM3436S KGADK2847O SRMUI5723N RDIXI7301N B
5 ADITC6567L HUOCT5660P AQCNE3753K FUMGY1428B D
6 BAEDP8491P IAGQG4816B TXXQH6337M SDACH5752D C我编写了这个循环,比较(col1,col2)和(col3,col4)的所有组合之间的不同字符串距离度量:
method = c("osa", "lv", "dl", "hamming", "lcs", "qgram", "cosine", "jaccard", "jw","soundex")
library(stringdist)
results = list()
for (i in 1:length(method))
{
method_i = method[i]
name_1_i = paste0("col1_col_2", method_i)
name_2_i = paste0("col3_col_4", method_i)
p1_i = stringdistmatrix(col1, col2, method = method_i, useNames = "string") %>%
as_tibble(rownames = "a") %>%
pivot_longer(-1, names_to = "b", values_to = name_1_i)
p2_i = stringdistmatrix(col3, col4, method = method_i, useNames = "string") %>%
as_tibble(rownames = "a") %>%
pivot_longer(-1, names_to = "b", values_to = name_2_i)
p1_i = p1_i[,3]
p2_i = p2_i[,3]
final_i = cbind(p1_i, p2_i)
results[[i]] = final_i
}
final = do.call(cbind.data.frame, results)
final = cbind(col1,col2, col3,col4, final)
average_col1_col2_dist = (final$col1_col_2osa + final$col1_col_2lv + final$col1_col_2dl + final$col1_col_2hamming + final$col1_col_2lcs + final$col1_col_2qgram + final$col1_col_2cosine + final$col1_col_2jaccard + final$col1_col_2jw + final$col1_col_2soundex)/10
average_col3_col4_dist = ( final$col3_col_4osa + final$col3_col_4lv + final$col3_col_4dl + final$col3_col_4hamming + final$col3_col_4lcs + final$col3_col_4qgram + final$col3_col_4cosine + final$col3_col_4jaccard + final$col3_col_4jw + final$col3_col_4soundex)/10
final = data.frame( col1, col2, col3, col4, average_col1_col2_dist, average_col3_col4_dist)
final = scale(final)现在,我想把它变成一个“双循环”,并进行相同的比较,但是应该只在每个“组”:中进行比较。
results = list()
for (i in 1:length(method))
for (j in 1:length(unique(example$group))
{
{
groups_j = unique(example$group[j])
my_data_i = file[which(file$fsa == groups_j ), ]
method_i = method[i]
name_1_i = paste0("col1_col_2", method_i)
name_2_i = paste0("col3_col_4", method_i)
p1_i = stringdistmatrix(my_data_i$col1, my_data_i$col2, method = method_i, useNames = "string") %>%
as_tibble(rownames = "a") %>%
pivot_longer(-1, names_to = "b", values_to = name_1_i)
p2_i = stringdistmatrix(my_data_i$col3, my_data_i$col4, method = method_i, useNames = "string") %>%
as_tibble(rownames = "a") %>%
pivot_longer(-1, names_to = "b", values_to = name_2_i)
p1_i = p1_i[,3]
p2_i = p2_i[,3]
final_i = cbind(p1_i, p2_i)
results[[i]] = final_i
}
}
final = do.call(cbind.data.frame, results)
final = cbind(col1,col2, col3,col4, final)
average_col1_col2_dist = (final$col1_col_2osa + final$col1_col_2lv + final$col1_col_2dl + final$col1_col_2hamming + final$col1_col_2lcs + final$col1_col_2qgram + final$col1_col_2cosine + final$col1_col_2jaccard + final$col1_col_2jw + final$col1_col_2soundex)/10
average_col3_col4_dist = ( final$col3_col_4osa + final$col3_col_4lv + final$col3_col_4dl + final$col3_col_4hamming + final$col3_col_4lcs + final$col3_col_4qgram + final$col3_col_4cosine + final$col3_col_4jaccard + final$col3_col_4jw + final$col3_col_4soundex)/10
final = data.frame( col1, col2, col3, col4, average_col1_col2_dist, average_col3_col4_dist)
final = scale(final)但是我不断地发现这个错误:
Error:
! Column 1 must be named.
Use .name_repair to specify repair.
Caused by error in `repaired_names()`:
! Names can't be empty.
x Empty name found at location 1.有人知道我怎么解决这个问题吗?
谢谢!
回答 1
Stack Overflow用户
发布于 2022-11-26 19:12:26
在试图理解您所做的工作的过程中,我偏离了您的原始代码。它的大部分并不一定有什么问题!
你的代码
至于你的分组代码..。
你开始的时候
for(this in that)
for(this in that)
{
{括号嵌套在for语句中的内容。你需要
for(this in that) {
for(this in that) {
# or this works
for(this in that)
{
for(this in that)
{当您指定您的for标准时,您使用整数。但是,您可以只使用字符串,如
for(i in method) { # i is a string
# versus
for(i in 1:length(method)) { # i is an integer当您编写嵌套的for语句时,忽略了一个结束括号。
for(j in 1:length(unique(example$group)) # end parentheses missing!
# should have been
for(j in 1:length(unique(example$group)))
# easier to see like this:
for(j in 1:length(
unique(
example$group
)
)
)你知道吗?您可以将RStudio设置为使用“彩虹括号”,这对于确保您不会错过结束括号或括号非常好。转到Tools -> Global Options -> Code (弹出式左菜单) ->显示(菜单弹出中的顶部菜单)&“彩虹括号”是列表中的最后一项。这就是我当前外观设置的样子:
提取组时,您选择了数据集行,而不是唯一值。
# this selects jth row, then looks for unique values
groups_j = unique(example$group[j])
# you need to get the unique values, then iterate
group_j = unique(example$group)[j]
# the j goes outside the call for unique()在这段代码中,您编写了file和file$fsa。我假设这相当于example和example$group,因为在file中没有任何东西。
所有这些代码行都是这样做的。请记住,group在数据框架中,但它本身也是环境中的一个对象。
my_data_i = example[which(example$group == group_j), ] # this would work
my_data_i <- filter(example, groups == group_j) # this would work
my_data_i <- example[group == group_j, ] # this would work
my_data_i <- example[example$group == group_j, ] # this would work如果您迭代组而不是索引,可以跳过group_j的创建,这是您使用j的唯一一次。
for(j in unique(example$group)) {
my_data_i <- example[example$group == j, ]
}当您单独使用i和i发送结果时,您将在j上用每次迭代覆盖数据。
第一组迭代可以在results[[i]]中进行,但下一个group可以绑定到该数据,也可以放在列表中的列表中。
例如:
results[[1]] <- group 1, method 1
results[[1]] <- rbind(results[[1]], [group 2, method 1])
# or
results[[1]][[1]] <- group 1, method 1
results[[1]][[2]] <- group 2, method 1考虑到列表的这两个选项(上面),第一个选项将允许您剩下的代码(创建最终代码、平均值等)。没有任何变化就能工作。但是,如果使用第二个选项(上面),则需要修改该代码。
如果您将for(j留给1:length,这将起作用:
if(j < 2) {
results[[i]] <- final_i
} else {
results[[i]] <- rbind(results[[i]], final_i)
}如果您使用for(j in unique(example$group)),您可以使用以下内容:
if(isTRUE(j == unique(example$group)[1])) { # isTRUE() to avoid null errors
results[[i]] <- final_i
} else {
results[[i]] <- rbind(results[[i]], final_i)
}嵌套的for语句都在一个块中。
results = list()
for (i in 1:length(method)) { # bracket missing here; it was in the wrong place
for (j in 1:length(unique(example$group))) { # missing a parentheses here
# { # this needs to be after each for statement
# groups_j = unique(example$group[j]) # you have selected the jth row, not the jth unique
group_j = unique(example$group)[j] # the selection goes outside the call for unique()
# use things like print or message to check what your function does
# print(group_j)
# message('this is a message ', group_j) # notice the different color in the console?
my_data_i <- example[group == group_j, ] # this would work
method_i = method[i]
name_1_i = paste0("col1_col_2", method_i)
name_2_i = paste0("col3_col_4", method_i)
p1_i = stringdistmatrix(my_data_i$col1, my_data_i$col2, method = method_i, useNames = "string") %>%
as_tibble(rownames = "a") %>%
pivot_longer(-1, names_to = "b", values_to = name_1_i)
p2_i = stringdistmatrix(my_data_i$col3, my_data_i$col4, method = method_i, useNames = "string") %>%
as_tibble(rownames = "a") %>%
pivot_longer(-1, names_to = "b", values_to = name_2_i)
p1_i = p1_i[,3]
p2_i = p2_i[,3]
final_i = cbind(p1_i, p2_i)
# results[[i]] = final_i # you replace this content everytime you change groups
# you need to append the values between groups (assuming you want one column per test type)
# first append, then combine
if(j < 2) { # use < instead of == to avoid null error
results[[i]] <- final_i
} else {
results[[i]] <- rbind(results[[i]], final_i)
}
}
}我的代码可以完成同样的任务
我增加了一些制衡,以使它更有活力。可以将任意数量的列、方法或组发送到grpComp。
它使用tidyverse、glue和stringdist。
第一个函数由另一个函数调用。
library(tidyverse)
library(stringdist)
library(glue)
strD <- function(c1, c2, mm) { # input column 1; column 2; measurement method
res <- stringdistmatrix(c1, c2, method = mm, useNames = 'string')
f_res <- matrix(res) # extract values and flatten
}这是按组函数计算的距离。
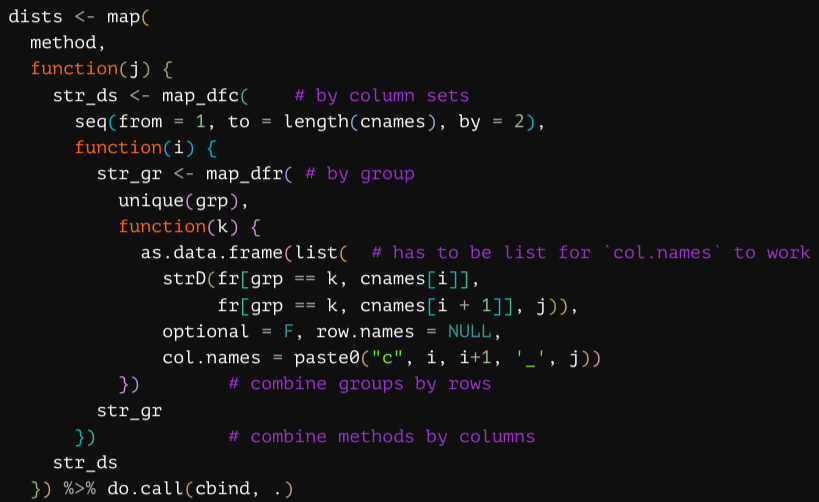
grpComp <- function(fr, methods, grp) { # data frame of columns to compare,
# methods to use, groups (vector same length as df rows)
cnames <- names(fr)
if(length(cnames) %% 2 != 0) {
message('there are an uneven number of columns to compare')
break # something's wrong
}
if(length(grp) != nrow(fr)) {
message('there groups vector length must match number of rows in the data')
break # something's wrong
}
# extract distances
dists <- map(
method,
function(j) {
str_ds <- map_dfc( # by column sets
seq(from = 1, to = length(cnames), by = 2),
function(i) {
str_gr <- map_dfr( # by group
unique(grp),
function(k) {
as.data.frame(list( # has to be list for `col.names` to work
strD(fr[grp == k, cnames[i]],
fr[grp == k, cnames[i + 1]], j)),
optional = F, row.names = NULL,
col.names = paste0("c", i, i+1, '_', j))
}) # combine groups by rows
str_gr
}) # combine methods by columns
str_ds
}) %>% do.call(cbind, .)
ncnames <- names(dists) %>% substr(1, 3) %>% unique() # determine unique col groups
for(m in ncnames) { # get averages for each comparison set
dists <- mutate(dists,
"ave_{m}" := rowMeans(across(contains(m))) %>% scale())
}
dists <- select(dists, contains('ave'))
}这就是你使用这段代码的方式。
test5 <- grpComp(example[, 1:4], methods, example$group)尽管您对非分组数据的功能是有效的,但我想我也会包括该代码。
strComp <- function(fr, methods) { # data frame of columns to compare, methods to use
cnames <- names(fr)
if(length(cnames) %% 2 != 0) {
message('there are an uneven number of columns to compare')
break # something's wrong
}
# extract distances
dists <- map(
method,
function(j) {
str_ds <- map_dfc(
seq(from = 1, to = length(cnames), by = 2),
function(i) {
as.data.frame(list( # has to be list for `col.names` to work
strD(fr[, cnames[i]], fr[, cnames[i + 1]], j)), optional = F,
col.names = paste0("c", i, i+1, '_', j))
})
str_ds
}) %>% do.call(cbind, .)
ncnames <- names(dists) %>% substr(1, 3) %>% unique() # determine unique col groups
for(k in ncnames) { # get averages for each comparison set
dists <- mutate(dists,
"ave_{k}" := rowMeans(across(contains(k))) %>% scale())
}
dists <- select(dists, contains('ave'))
}要使用此功能:
test4 <- strComp(example[, 1:4], methods)https://stackoverflow.com/questions/74557454
复制相似问题

