利用GEKKO生物反应器模拟乙醇生产
利用GEKKO生物反应器模拟乙醇生产
提问于 2022-11-08 14:42:50
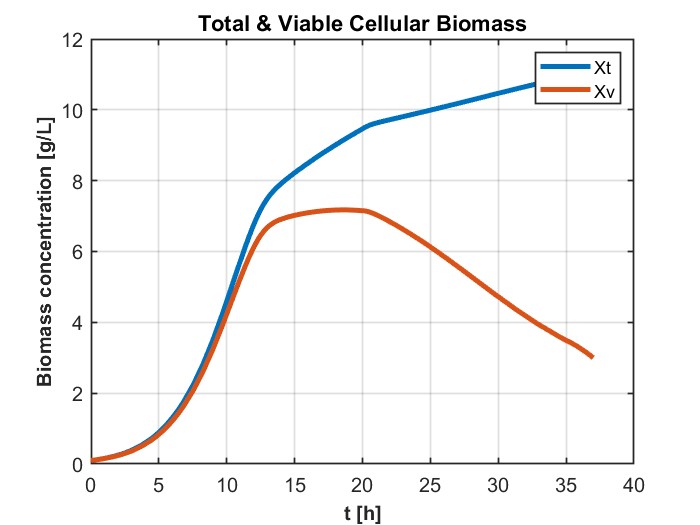
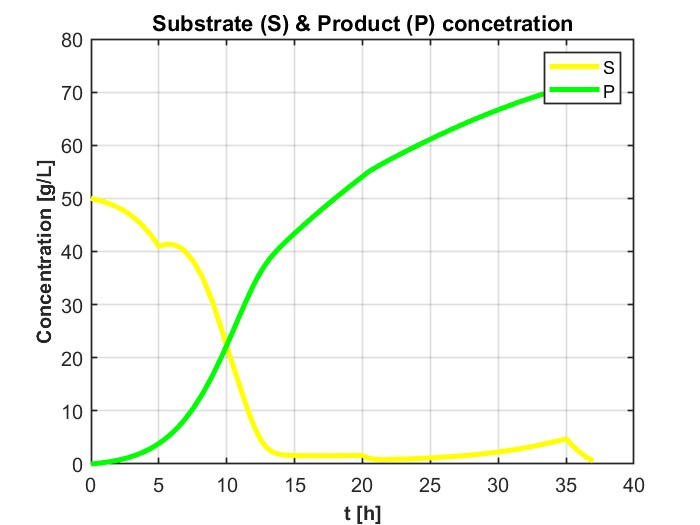
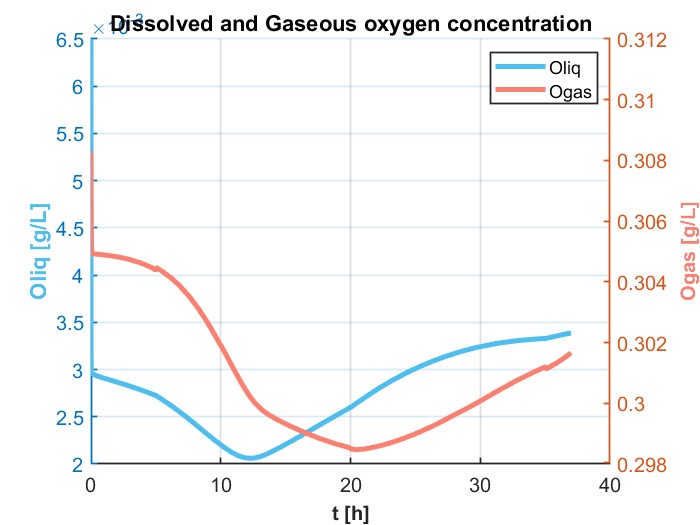
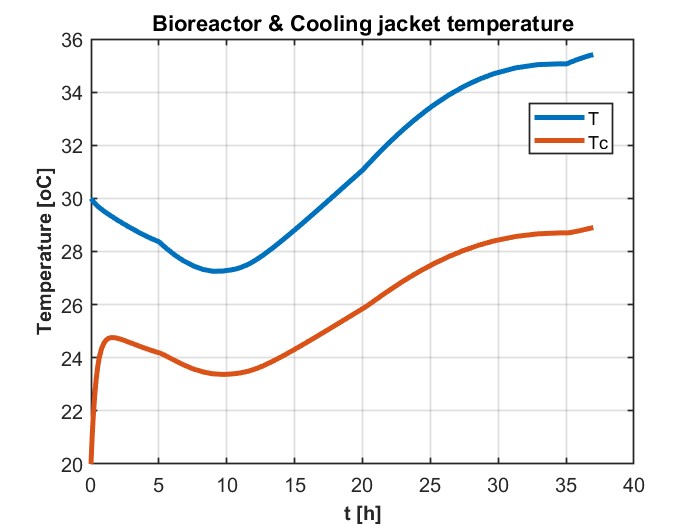
我正在尝试模拟一个DAE系统,它解决了用GEKKO生产乙醇的进料批量生物反应器的问题。这是为了以后我可以更容易地优化它,以最大限度地生产乙醇。它以前是在MATLAB中解决的,得到了如下数字所示的结果:
,
,
,
,
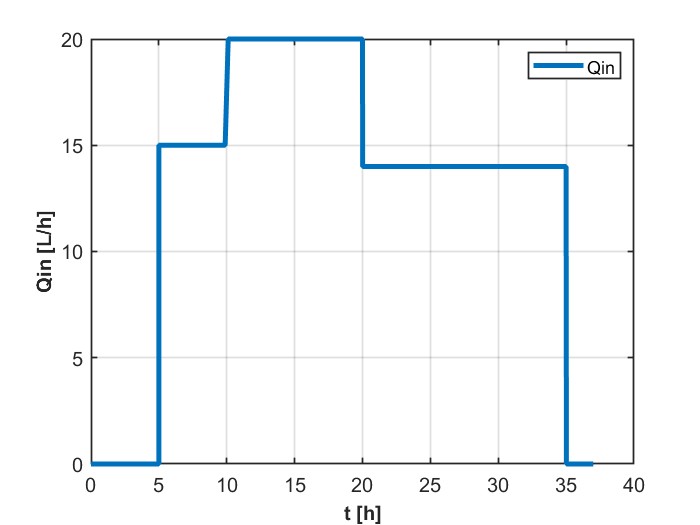
我现在的问题是,我不能用GEKKO产生相同的结果,给出所有相同的常量和变量的值。无法找到解决方案,但收敛的时间较短,例如: m.time= np.linspace(0、1、11)。知道我的代码出了什么问题吗?
需要解决的最初制度是:
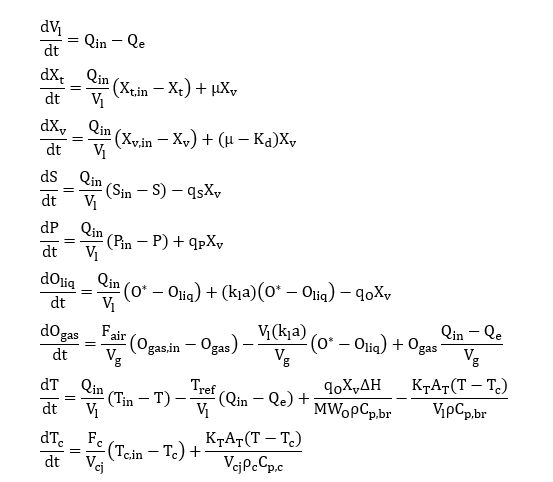


from gekko import GEKKO
import numpy as np
import matplotlib.pyplot as plt
m = GEKKO(remote=False)
# Create time vector: t=[0, 0.1, 0.2,...,36.9,37], [hours]
nt = 371
m.time = np.linspace(0,37,nt)
# Define constants and parameters
#################################
# Kinetic Parameters
a1 = m.Const(value=0.05, name='a1') # Ratkowsky parameter [oC-1 h-0.5]
aP = m.Const(value=4.50, name='aP') # Growth-associated parameter for EtOh production [-]
AP1 = m.Const(value=6.0, name='AP1') # Activation energy parameter for EtOh production [oC]
AP2 = m.Const(value=20.3, name='AP2') # Activation energy parameter for EtOh production [oC]
b1 = m.Const(value=0.035, name='b1') # Parameter in the exponential expression of the maximum specific growth rate expression [oC-1]
b2 = m.Const(value=0.15, name='b2') # Parameter in the exponential expression of the maximum specific growth rate expression [oC-1]
b3 = m.Const(value=0.40, name='b3') # Parameter in the exponential expression of the specific death rate expression [oC-1]
c1 = m.Const(value=0.38, name='c1') # Constant decoupling factor for EtOh [gP gX-1 h-1]
c2 = m.Const(value=0.29, name='c2') # Constant decoupling factor for EtOh [gP gX-1 h-1]
k1 = m.Const(value=3, name='k1') # Parameter in the maximum specific growth rate expression [oC]
k2 = m.Const(value=55, name='k2') # Parameter in the maximum specific growth rate expression [oC]
k3 = m.Const(value=60, name='k3') # Parameter in the growth-inhibitory EtOH concentration expression [oC]
k4 = m.Const(value=50, name='k4') # Temperature at the inflection point of the specific death rate sigmoid curve [oC]
Pmaxb = m.Const(value=90, name='Pmaxb') # Temperature-independent product inhibition constant [g L-1]
PmaxT = m.Const(value=90, name='PmaxT') # Maximum value of product inhibition constant due to temperature [g L-1]
Kdb = m.Const(value=0.025, name='Kdb') # Basal specific cellular biomass death rate [h-1]
KdT = m.Const(value=30, name='KdT') # Maximum value of specific cellular biomass death rate due to temperature [h-1]
KSX = m.Const(value=5, name='KSX') # Glucose saturation constant for the specific growth rate [g L-1]
KOX = m.Const(value=0.0005, name='KOX') # Oxygen saturation constant for the specific growth rate [g L-1]
qOmax = m.Const(value=0.05, name='qOmax') # Maximum specific oxygen consumption rate [h-1]
# Metabolic Parameters
YPS = m.Const(value=0.51, name='YPS') # Theoretical yield of EtOH on glucose [gP gS-1]
YXO = m.Const(value=0.97, name='YXO') # Theoretical yield of biomass on oxygen [gX gO-1]
YXS = m.Const(value=0.53, name='YXS') # Theoretical yield of biomass on glucose [gX gS-1]
# Physicochemical and thermodynamic parameters
Chbr = m.Const(value=4.18, name='Chbr') # Heat capacity of the mass of reaction [J g-1 oC-1]
Chc = m.Const(value=4.18, name='Chc') # Heat capacity of cooling agent [J g-1 oC-1]
deltaH = m.Const(value=518.e3, name='deltaH') # Heat of reaction of fermentation [J mol-1 O2]
Tref = m.Const(value=25, name='Tref') # Reference temperature [oC]
KH = m.Const(value=200, name='KH') # Henry's constant for oxygen in the fermentation broth [atm L mol-1]
z = m.Const(value=0.792, name='z') # Oxygen compressibility factor [-]
R = m.Const(value=0.082, name='R') # Ideal gas constant [L atm mol-1 oC-1]
kla0 = m.Const(value=100, name='kla0') # Temperature-independent volumetric oxygen transfer coefficient [-h]
KT = m.Const(value=36.e4, name='KT') # Heat transfer coefficient [J h-1 m-2 oC-1]
rho = m.Const(value=1080, name='rho') # Density of the fermentation broth [g L-1]
rhoc = m.Const(value=1000, name='rhoc') # Density of the cooling agent [g L-1]
MO = m.Const(value=15.999, name='MO') # Molecular weight of oxygen [g mol-1]
# Bioreactor design data
AT = m.Const(value=1, name='AT') # Bioreactor heat transfer area [m2]
V = m.Const(value=2000, name='V') # Bioreactor working volume [L]
Vcj = m.Const(value=250, name='Vcj') # Cooling jacket volume [L]
Ogasin = m.Const(value=0.305, name='Ogasin') # Oxygen concentration in airflow inlet [g L-1]
# Define variables
##################
mi = m.Var(name='mi')
# I want Qin to be a step function: Qin = Qin0 + 15H(t-5) + 5H(t-10) - 6H(t-20) - 14H(t-35), where H(t-t0) heaviside function
Qin_step = np.zeros(nt)
Qin_step[50:101] = 15
Qin_step[101:201] = 20
Qin_step[201:350] = 14
Qin = m.Param(value=Qin_step, name='Qin')
# Fixed variables, they are constant throughout the time horizon
Xtin = m.FV(value=0, name='Xtin')
Xvin = m.FV(value=0, name='Xvin')
Qe = m.FV(value=0, name='Qe')
Sin = m.FV(value=400, lb=0, ub=1500)
Pin = m.FV(value=0, name='Pin')
Fc = m.FV(value=40, name='Fc')
Fair = m.FV(value=60000, name='Fair')
Tin = m.FV(value=30, name='Tin')
Tcin = m.FV(value=15, name='Tcin')
Vl = m.Var(value=1000, lb=-0.0, ub=0.75*V, name='Vl')
Xt = m.Var(value=0.1, lb=-0.0, ub=10, name='Xt')
Xv = m.Var(value=0.1, lb=-0.0, ub=10, name='Xv')
S = m.Var(value=400, lb=+0.0, ub=10000, name='S')
P = m.Var(value=0, name='P')
Ol = m.Var(value=0.0065, name= 'Ol')
Og = m.Var(value=0.305, name='Og')
T = m.Var(value=30, lb=20, ub=40, name='T')
Tc = m.Var(value=20, lb=0, ub=30, name='Tc')
Sf_cum = m.Var(value=0, name='Sf_cum')
t = m.Var(value=0, name='Time')
# Define algebraic equations
############################
# Specific growth rate of cell mass
mimax = m.Intermediate(((a1*(T - k1))*(1 - m.exp(b1 * (T - k2)) )) ** 2)
Pmax = m.Intermediate(Pmaxb + PmaxT/(1- m.exp(-b2*(T-k3))))
m.Equation(mi == mimax * (S / (KSX + S)) * (Ol / (KOX + Ol)) * (1 - P / Pmax) * (1 / (1 + m.exp(-(100 - S)))))
mi = m.if3(condition=mi, x1=0, x2=mi)
# Specific production rate of EtOH
bP = m.if3(condition=S, x1=0, x2=c1*m.exp(-AP1/T) - c2*m.exp(-AP2/T))
qP = m.Intermediate(aP*mi + bP)
# Specific consumption rate of glucose
qS = m.Intermediate(mi/YXS + qP/YPS)
# Specific consumption rate of oxygen
qO = m.Intermediate(qOmax*Ol/YXO/(KOX+Ol))
# Specific biological deactivation rate of cell mass
Kd = m.Intermediate(Kdb + KdT/(1+m.exp(-b3*(T-k4))))
# Saturation concentration of oxygen in culture media
Ostar = m.Intermediate(z*Og*R*T/KH)
# Oxygen mass transfer coefficient
kla = m.Intermediate(kla0*1.2**(T-20))
# Bioreactor phases equation
Vg = m.Intermediate(V - Vl)
# Define differential equations
###############################
m.Equation(Vl.dt() == Qin - Qe)
m.Equation(Xt.dt() == Qin/Vl*(Xtin-Xt) + mi*Xv)
m.Equation(Xv.dt() == Qin/Vl*(Xvin-Xv) + Xv*(mi-Kd))
m.Equation(S.dt() == Qin/Vl*(Sin-S) - qS*Xv)
m.Equation(P.dt() == Qin/Vl*(Pin - P) + qP*Xv)
m.Equation(Ol.dt() == Qin/Vl*(Ostar-Ol) + kla*(Ostar-Ol) - qO*Xv)
m.Equation(Og.dt() == Fair/Vg*(Ogasin-Og) - Vl*kla/Vg*(Ostar-Ol) + Og*(Qin-Qe)/Vg)
m.Equation(T.dt() == Qin/Vl*(Tin-T) - Tref/Vl*(Qin-Qe) + qO*Xv*deltaH/MO/rho/Chbr - KT*AT*(T-Tc)/Vl/rho/Chbr)
m.Equation(Tc.dt() == Fc/Vcj*(Tcin - Tc) + KT*AT*(T-Tc)/Vcj/rhoc/Chc)
m.Equation(Sf_cum.dt() == Qin*Sin)
m.Equation(t.dt() == 1)
# solve ODE
m.options.IMODE = 6
# m.open_folder()
m.solve(display=True)
# Plot results
plt.figure(1)
plt.title('Total & Viable Cellular Biomass')
plt.plot(m.time, Xv.value, label='Xv')
plt.plot(m.time, Xt.value, label='Xt')
plt.legend()
plt.ylabel('Biomass concentration [g/L]')
plt.xlabel('Time [h]')
plt.grid()
plt.minorticks_on()
plt.ylim(0)
plt.xlim(m.time[0],m.time[-1])
plt.tight_layout()
plt.figure(2)
plt.title('Substrate (S) & Product (P) concentration')
plt.plot(m.time, S.value, label='S')
plt.plot(m.time, P.value, label='P')
plt.legend()
plt.ylabel('Concentration [g/L]')
plt.xlabel('Time [h]')
plt.grid()
plt.minorticks_on()
plt.ylim(0)
plt.xlim(m.time[0],m.time[-1])
plt.tight_layout()
plt.figure(3)
plt.title('Bioreactor & Cooling jacket temperature')
plt.plot(m.time, T.value, label='T')
plt.plot(m.time, Tc.value, label='Tc')
plt.legend()
plt.ylabel('Temperature [oC]')
plt.xlabel('Time [h]')
plt.grid()
plt.minorticks_on()
plt.ylim(0)
plt.xlim(m.time[0],m.time[-1])
plt.tight_layout()
fig4, ax = plt.subplots()
ax.title.set_text('Dissolved & Gaseous Oxygen concentration')
lns1 = ax.plot(m.time, Ol.value, label='[Oliq]', color='c')
ax.set_xlabel('Time [h]')
ax.set_ylabel('Oliq [g/L]', color='c')
ax.minorticks_on()
ax2 = ax.twinx()
lns2 = ax2.plot(m.time, Og.value, label='[Ogas]', color='y')
ax2.set_ylabel('Ogas [g/L]', color='y')
ax2.minorticks_on()
lns = lns1 + lns2
labs = [l.get_label() for l in lns]
ax.legend(lns, labs, loc='best')
ax.grid()
fig4.tight_layout()
plt.figure(4)
plt.figure(5)
plt.title('Feeding Policy')
plt.plot(m.time, Qin.value, label='Qin')
plt.legend()
plt.ylabel('Qin [L/h]')
plt.xlabel('Time [h]')
plt.grid()
plt.minorticks_on()
plt.ylim(0)
plt.xlim(m.time[0],m.time[-1])
plt.tight_layout()
plt.show()回答 1
Stack Overflow用户
回答已采纳
发布于 2022-11-09 02:51:40
很好的申请!以下是一些改进收敛性的建议。
- 在模拟时移除上下界。这导致了“找不到解决方案”错误。
Vl = m.Var(value=1000, name='Vl') # lb=-0.0, ub=0.75*V
Xt = m.Var(value=0.1, name='Xt') # lb=-0.0, ub=10
Xv = m.Var(value=0.1, name='Xv') # lb=-0.0, ub=10
S = m.Var(value=400, name='S') # lb=+0.0, ub=10000
P = m.Var(value=0, name='P')
Ol = m.Var(value=0.0065, name= 'Ol')
Og = m.Var(value=0.305, name='Og')
T = m.Var(value=30, name='T') # lb=20, ub=40
Tc = m.Var(value=20, name='Tc') # lb=0, ub=30- 重新安排方程,以避免除以零(如有可能)。对于大多数方程,体积项可以移动到方程的左边,以避免分母中的变量。
m.Equation(Vl.dt() == Qin - Qe)
m.Equation(Vl*Xt.dt() == Qin*(Xtin-Xt) + mi*Vl*Xv)
m.Equation(Vl*Xv.dt() == Qin*(Xvin-Xv) + Xv*Vl*(mi-Kd))
m.Equation(Vl*S.dt() == Qin*(Sin-S) - qS*Vl*Xv)
m.Equation(Vl*P.dt() == Qin*(Pin - P) + qP*Vl*Xv)
m.Equation(Vl*Ol.dt() == Qin*(Ostar-Ol) + Vl*kla*(Ostar-Ol) - qO*Vl*Xv)
m.Equation(Vg*Og.dt() == Fair*(Ogasin-Og) - Vl*kla*(Ostar-Ol) + Og*(Qin-Qe))
m.Equation(Vl*T.dt() == Qin*(Tin-T) - Tref*(Qin-Qe) \
+ Vl*qO*Xv*deltaH/MO/rho/Chbr - KT*AT*(T-Tc)/rho/Chbr)
m.Equation(Vcj*Tc.dt() == Fc*(Tcin - Tc) + KT*AT*(T-Tc)/rhoc/Chc)
m.Equation(Sf_cum.dt() == Qin*Sin)- 使用
APOPT解算器提高速度,增加NODES=3以提高精度。IMODE=7是一种在零自由度情况下提高求解速度的顺序仿真(仿真,#equations=#variables)。
m.options.SOLVER= 1
m.options.IMODE = 7
m.options.NODES = 3 - 下面是一种基于时间的步骤输入的更简单的定义方法。
# I want Qin to be a step function:
# Qin = Qin0 + 15H(t-5) + 5H(t-10) - 6H(t-20) - 14H(t-35)
# where H(t-t0) heaviside function
Qin_step = np.zeros(nt)
Qin_step[np.where(tm>=5)] += 15
Qin_step[np.where(tm>=10)] += 5
Qin_step[np.where(tm>=20)] -= 6
Qin_step[np.where(tm>=35)] -= 14
Qin = m.Param(value=Qin_step, name='Qin')- 避免
if3()(如果可能),并在变量定义中用下界替换:
#mi = m.if3(condition=mi, x1=0, x2=mi)
mi = m.Var(name='mi',lb=0)
m.Equation(mi == mimax * (S / (KSX+S)) * (Ol/(KOX + Ol)) \
* (1 - P/Pmax) * (1 / (1+m.exp(-(100-S)))))或者松弛变量 slk,它避免了m.if3()引入的二进制交换变量。
slk = m.Var(0,lb=0)
mi_u = m.Var(name='mi_u')
mi = m.Var(name='mi',lb=0)
m.Equation(mi = mi_u + slk)
m.Minimize(slk)- (可选)如果出现收敛问题,则在开始时插入一些额外的小步骤。这在以后开始优化时非常有用。
# Create time vector: t=[0, 0.1, 0.2,...,36.9,37], [hours]
tm = np.linspace(0,37,371)
# Insert smaller time steps at the beginning
tm = np.insert(tm,1,[0.001,0.005,0.01,0.05])- (稍后)当您需要优化时,当您需要对优化问题进行初步猜测时,收敛到模拟中通常是很有帮助的。
m.options.IMODE=7
m.solve()
m.options.IMODE=6
m.solve()这是完整的剧本。
from gekko import GEKKO
import numpy as np
import matplotlib.pyplot as plt
m = GEKKO(remote=False)
# Create time vector: t=[0, 0.1, 0.2,...,36.9,37], [hours]
tm = np.linspace(0,37,371)
# Insert smaller time steps at the beginning
tm = np.insert(tm,1,[0.001,0.005,0.01,0.05])
m.time = tm
nt = len(tm)
# Define constants and parameters
#################################
# Kinetic Parameters
a1 = m.Const(value=0.05, name='a1') # Ratkowsky parameter [oC-1 h-0.5]
aP = m.Const(value=4.50, name='aP') # Growth-associated parameter for EtOh production [-]
AP1 = m.Const(value=6.0, name='AP1') # Activation energy parameter for EtOh production [oC]
AP2 = m.Const(value=20.3, name='AP2') # Activation energy parameter for EtOh production [oC]
b1 = m.Const(value=0.035, name='b1') # Parameter in the exponential expression of the maximum specific growth rate expression [oC-1]
b2 = m.Const(value=0.15, name='b2') # Parameter in the exponential expression of the maximum specific growth rate expression [oC-1]
b3 = m.Const(value=0.40, name='b3') # Parameter in the exponential expression of the specific death rate expression [oC-1]
c1 = m.Const(value=0.38, name='c1') # Constant decoupling factor for EtOh [gP gX-1 h-1]
c2 = m.Const(value=0.29, name='c2') # Constant decoupling factor for EtOh [gP gX-1 h-1]
k1 = m.Const(value=3, name='k1') # Parameter in the maximum specific growth rate expression [oC]
k2 = m.Const(value=55, name='k2') # Parameter in the maximum specific growth rate expression [oC]
k3 = m.Const(value=60, name='k3') # Parameter in the growth-inhibitory EtOH concentration expression [oC]
k4 = m.Const(value=50, name='k4') # Temperature at the inflection point of the specific death rate sigmoid curve [oC]
Pmaxb = m.Const(value=90, name='Pmaxb') # Temperature-independent product inhibition constant [g L-1]
PmaxT = m.Const(value=90, name='PmaxT') # Maximum value of product inhibition constant due to temperature [g L-1]
Kdb = m.Const(value=0.025, name='Kdb') # Basal specific cellular biomass death rate [h-1]
KdT = m.Const(value=30, name='KdT') # Maximum value of specific cellular biomass death rate due to temperature [h-1]
KSX = m.Const(value=5, name='KSX') # Glucose saturation constant for the specific growth rate [g L-1]
KOX = m.Const(value=0.0005, name='KOX') # Oxygen saturation constant for the specific growth rate [g L-1]
qOmax = m.Const(value=0.05, name='qOmax') # Maximum specific oxygen consumption rate [h-1]
# Metabolic Parameters
YPS = m.Const(value=0.51, name='YPS') # Theoretical yield of EtOH on glucose [gP gS-1]
YXO = m.Const(value=0.97, name='YXO') # Theoretical yield of biomass on oxygen [gX gO-1]
YXS = m.Const(value=0.53, name='YXS') # Theoretical yield of biomass on glucose [gX gS-1]
# Physicochemical and thermodynamic parameters
Chbr = m.Const(value=4.18, name='Chbr') # Heat capacity of the mass of reaction [J g-1 oC-1]
Chc = m.Const(value=4.18, name='Chc') # Heat capacity of cooling agent [J g-1 oC-1]
deltaH = m.Const(value=518000, name='deltaH') # Heat of reaction of fermentation [J mol-1 O2]
Tref = m.Const(value=20, name='Tref') # Reference temperature [oC]
KH = m.Const(value=200, name='KH') # Henry's constant for oxygen in the fermentation broth [atm L mol-1]
z = m.Const(value=0.792, name='z') # Oxygen compressibility factor [-]
R = m.Const(value=0.082, name='R') # Ideal gas constant [L atm mol-1 oC-1]
kla0 = m.Const(value=100, name='kla0') # Temperature-independent volumetric oxygen transfer coefficient [-h]
KT = m.Const(value=360000, name='KT') # Heat transfer coefficient [J h-1 m-2 oC-1]
rho = m.Const(value=1080, name='rho') # Density of the fermentation broth [g L-1]
rhoc = m.Const(value=1000, name='rhoc') # Density of the cooling agent [g L-1]
MO = m.Const(value=32.0, name='MO') # Molecular weight of oxygen [g mol-1]
# Bioreactor design data
AT = m.Const(value=1, name='AT') # Bioreactor heat transfer area [m2]
V = m.Const(value=1800, name='V') # Bioreactor working volume [L]
Vcj = m.Const(value=50, name='Vcj') # Cooling jacket volume [L]
Ogasin = m.Const(value=0.305, name='Ogasin') # Oxygen concentration in airflow inlet [g L-1]
# Define variables
##################
mi = m.Var(name='mi',lb=0)
# I want Qin to be a step function:
# Qin = Qin0 + 15H(t-5) + 5H(t-10) - 6H(t-20) - 14H(t-35)
# where H(t-t0) heaviside function
Qin_step = np.zeros(nt)
Qin_step[np.where(tm>=5)] += 15
Qin_step[np.where(tm>=10)] += 5
Qin_step[np.where(tm>=20)] -= 6
Qin_step[np.where(tm>=35)] -= 14
Qin = m.Param(value=Qin_step, name='Qin')
# Fixed variables, they are constant throughout the time horizon
Xtin = m.FV(value=0, name='Xtin')
Xvin = m.FV(value=0, name='Xvin')
Qe = m.FV(value=0, name='Qe')
Sin = m.FV(value=400, lb=0, ub=1500)
Pin = m.FV(value=0, name='Pin')
Fc = m.FV(value=40, name='Fc')
Fair = m.FV(value=60000, name='Fair')
Tin = m.FV(value=30, name='Tin')
Tcin = m.FV(value=15, name='Tcin')
Vl = m.Var(value=1000, name='Vl') # lb=-0.0, ub=0.75*V
Xt = m.Var(value=0.1, name='Xt') # lb=-0.0, ub=10
Xv = m.Var(value=0.1, name='Xv') # lb=-0.0, ub=10
S = m.Var(value=50, name='S') # lb=+0.0, ub=10000
P = m.Var(value=0, name='P')
Ol = m.Var(value=0.0065, name= 'Ol')
Og = m.Var(value=0.305, name='Og')
T = m.Var(value=30, name='T') # lb=20, ub=40
Tc = m.Var(value=20, name='Tc') # lb=0, ub=30
Sf_cum = m.Var(value=0, name='Sf_cum')
#t = m.Var(value=0, name='Time')
# Define algebraic equations
############################
# Specific growth rate of cell mass
mimax = m.Intermediate(((a1*(T-k1))*(1-m.exp(b1*(T-k2))))** 2)
Pmax = m.Intermediate(Pmaxb + PmaxT/(1-m.exp(-b2*(T-k3))))
m.Equation(mi == mimax * (S / (KSX+S)) * (Ol/(KOX + Ol)) \
* (1 - P/Pmax) * (1 / (1+m.exp(-(100-S)))))
#mi = m.if3(condition=mi, x1=0, x2=mi)
# Specific production rate of EtOH
#bP = m.if3(condition=S, x1=0, x2=c1*m.exp(-AP1/T) - c2*m.exp(-AP2/T))
bP = m.Intermediate(c1*m.exp(-AP1/T) - c2*m.exp(-AP2/T))
qP = m.Intermediate(aP*mi + bP)
# Specific consumption rate of glucose
qS = m.Intermediate(mi/YXS + qP/YPS)
# Specific consumption rate of oxygen
qO = m.Intermediate(qOmax*Ol/YXO/(KOX+Ol))
# Specific biological deactivation rate of cell mass
Kd = m.Intermediate(Kdb + KdT/(1+m.exp(-b3*(T-k4))))
# Saturation concentration of oxygen in culture media
Ostar = m.Intermediate(z*Og*R*T/KH)
# Oxygen mass transfer coefficient
kla = m.Intermediate(kla0*1.2**(T-20))
# Bioreactor phases equation
Vg = m.Intermediate(V - Vl)
# Define differential equations
###############################
m.Equation(Vl.dt() == Qin - Qe)
m.Equation(Vl*Xt.dt() == Qin*(Xtin-Xt) + mi*Vl*Xv)
m.Equation(Vl*Xv.dt() == Qin*(Xvin-Xv) + Xv*Vl*(mi-Kd))
m.Equation(Vl*S.dt() == Qin*(Sin-S) - qS*Vl*Xv)
m.Equation(Vl*P.dt() == Qin*(Pin - P) + qP*Vl*Xv)
m.Equation(Vl*Ol.dt() == Qin*(Ostar-Ol) + Vl*kla*(Ostar-Ol) - qO*Vl*Xv)
m.Equation(Vg*Og.dt() == Fair*(Ogasin-Og) - Vl*kla*(Ostar-Ol) + Og*(Qin-Qe))
m.Equation(Vl*T.dt() == Qin*(Tin-T) - Tref*(Qin-Qe) \
+ Vl*qO*Xv*deltaH/MO/rho/Chbr - KT*AT*(T-Tc)/rho/Chbr)
m.Equation(Vcj*Tc.dt() == Fc*(Tcin - Tc) + KT*AT*(T-Tc)/rhoc/Chc)
m.Equation(Sf_cum.dt() == Qin*Sin)
#m.Equation(t.dt() == 1)
# solve ODE
m.options.SOLVER= 1
m.options.IMODE = 7
m.options.NODES = 3
# m.open_folder()
m.solve(disp=False)
# Plot results
plt.figure(1)
plt.title('Total & Viable Cellular Biomass')
plt.plot(m.time, Xv.value, label='Xv')
plt.plot(m.time, Xt.value, label='Xt')
plt.legend()
plt.ylabel('Biomass concentration [g/L]')
plt.xlabel('Time [h]')
plt.grid()
plt.minorticks_on()
plt.ylim(0)
plt.xlim(m.time[0],m.time[-1])
plt.tight_layout()
plt.figure(2)
plt.title('Substrate (S) & Product (P) concentration')
plt.subplot(2,1,1)
plt.plot(m.time, S.value, label='S')
plt.legend(); plt.grid()
plt.ylabel('Conc [g/L]')
plt.subplot(2,1,2)
plt.plot(m.time, P.value, label='P')
plt.legend(); plt.grid()
plt.ylabel('Conc [g/L]')
plt.xlabel('Time [h]')
plt.minorticks_on()
plt.ylim(0)
plt.xlim(m.time[0],m.time[-1])
plt.tight_layout()
plt.figure(3)
plt.title('Bioreactor & Cooling jacket temperature')
plt.plot(m.time, T.value, label='T')
plt.plot(m.time, Tc.value, label='Tc')
plt.legend()
plt.ylabel('Temperature [oC]')
plt.xlabel('Time [h]')
plt.grid()
plt.minorticks_on()
plt.ylim(0)
plt.xlim(m.time[0],m.time[-1])
plt.tight_layout()
fig4, ax = plt.subplots()
ax.title.set_text('Dissolved & Gaseous Oxygen concentration')
lns1 = ax.plot(m.time, Ol.value, label='[Oliq]', color='c')
ax.set_xlabel('Time [h]')
ax.set_ylabel('Oliq [g/L]', color='c')
ax.minorticks_on()
ax2 = ax.twinx()
lns2 = ax2.plot(m.time, Og.value, label='[Ogas]', color='y')
ax2.set_ylabel('Ogas [g/L]', color='y')
ax2.minorticks_on()
lns = lns1 + lns2
labs = [l.get_label() for l in lns]
ax.legend(lns, labs, loc='best')
ax.grid()
fig4.tight_layout()
plt.figure(4)
plt.figure(5)
plt.title('Feeding Policy')
plt.plot(m.time, Qin.value, label='Qin')
plt.legend()
plt.ylabel('Qin [L/h]')
plt.xlabel('Time [h]')
plt.grid()
plt.minorticks_on()
plt.ylim(0)
plt.xlim(m.time[0],m.time[-1])
plt.tight_layout()
plt.figure(6)
plt.title('Check >=0 Constraints')
plt.subplot(2,1,1)
plt.plot(tm,bP.value,label='bP')
plt.legend(); plt.grid()
plt.subplot(2,1,2)
plt.plot(tm,mi.value,label='mi')
plt.legend(); plt.grid()
plt.show()这些图用更新的参数复制Matlab图(谢谢建议)。如果我们能帮助进一步的问题,请告诉我们。
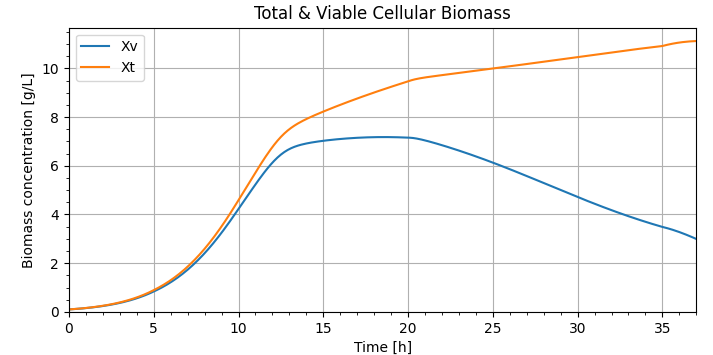
Python版本
下面是一个等效的Python版本。如果仍然需要,我创建它是为了帮助进行等效性测试。
import numpy as np
from scipy.integrate import odeint
from scipy.interpolate import interp1d
import matplotlib.pyplot as plt
#Simulate fed-batch operation
# Specify the simulation time (hrs)
tspan = np.linspace(0,37,371); t=tspan
# Specify values of control variables [Qin0 Xtin Xvin Qe Sin Fc Fair Tin Tcin]
u0 = [0.0,0.0,0.0,0.0,0.0,400,40,60000,30,15]
# Specify initial conditions [Xt Xv S P Oliq Ogas T Tc Vl Gloss MP]
x0 = [0.1,0.1,50,0,0.0065,0.305,30,20,1000,0.0,0.0]
ux0 = tuple(u0 + x0)
#Qin = 15*heaviside(t-5) + 5*heaviside(t-10)
# - 6*heaviside(t-20) - 14*heaviside(t-35)
Qin = np.zeros_like(tspan)
Qin[np.where(tspan>=5)] += 15
Qin[np.where(tspan>=10)] += 5
Qin[np.where(tspan>=20)] -= 6
Qin[np.where(tspan>=35)] -= 14
QinInterp = interp1d(tspan,Qin,bounds_error=False)
def ethanol(x,t,Qin0,Qin,Xtin,Xvin,Qe,Sin,Fc,Fair,
Tin,Tcin,Xt0,Xv0,S0,P0,Oliq0,Ogas0,T0,
Tc0,Vl0,Sf_cum0,Time0):
## #Initial Conditions
## Xt0 = u[10] # Initial total cellular biomass, [g L-1]
## Xv0 = u[11] # Initial viable cellular biomass, [g L-1]
## S0 = u[12] # Initial substrate/Glucose concentration, [g L-1]
## P0 = u[13] # Initial product/Ethanol concentration, [g L-1]
## Oliq0 = u[14] # Initial Dissolved oxygen concentration, [g L-1]
## Ogas0 = u[15] # Initial Gas phase oxygen (bubbles) in the fermentation broth, [g L-1]
## T0 = u[16] # Initial Temperature in the bioreactor, [oC]
## Tc0 = u[17] # Initial Temperature of the cooling agent in the jacket, [oC]
## Vl0 = u[18] # Initial Culture volume in the bioreactor, [L]
## Sf_cum0 = u[19] # Initial Cumulative substrate/glucose fed to the bioreactor, [g]
## Time0 = u[20] # Initial batch time, [h]
##
## #Control variables
## Qin0 = u[0] # Volumetric inflow rate, [l/h-1]
## Qin = u[1] # Volumetric inflow rate, [l/h-1]
## Xtin = u[2] # Total biomass concentration in the bioreactor feed, [g L-1]
## Xvin = u[3] # Viable biomass concentration in the bioreactor feed, [g L-1]
## Qe = u[4] # Volumetric outflow rate, [l/h-1]
## Sin = u[5] # Substrate/Glucose concentration in bioreactor feed, [g L-1]
## Fc = u[6] # Cooling agent inlet volumetric flowrate, [L h-1]
## Fair = u[7] # Airflow inlet volumetric flowrate, [L h-1]
## Tin = u[8] # Temperature of bioreactor feed, [oC]
## Tcin = u[9] # Temperature of cooling agent inlet, [oC]
# 1D Interpolation for Qin
Qin = QinInterp(t)
#Definition of model parameters
#Kinetic parameters
a1 = 0.05 # Ratkowsky parameter [oC-1 h-0.5]
aP = 4.50 # Growth-associated parameter for ethanol production, [-]
AP1 = 6.0 # Activation energy parameter for ethanol production, [oC]
AP2 = 20.3 # Activation energy parameter for ethanol production, [oC]
b1 = 0.035 # Parameter in the exponential expression of the maximum specific growth rate np.expression, [oC-1]
b2 = 0.15 # Parameter in the exponential expression of the growth inhibitory ethanol concentration np.expression, [oC-1]
b3 = 0.40 # Parameter in the exponential np.expression of the specific death rate expression,[oC-1]
c1 = 0.38 # Constant decoupling factor for ethanol production, [gP gX-1 h-1]
c2 = 0.29 # Constant decoupling factor for ethanol production, [gP gX-1 h-1]
k1 = 3.00 # Parameter in the maximum specific growth rate expression, [oC]
k2 = 55.0 # Parameter in the maximum specific growth rate expression, [oC]
k3 = 60.0 # Parameter in the growth-inhibitory ethanol concentration expression, [oC]
k4 = 50.0 # Temperature at the inflection point of the specific death rate sigmoid curve, [oC]
Pmaxb = 90 # Temperature-independent product inhibition constant, [g L-1]
PmaxT = 90 # Maximum value of product inhibition constant due to temperature, [g L-1]
Kdb = 0.025 # Basal specific cellular biomass death rate, [h-1]
KdT = 30.00 # Maximum value of specific cellular biomass death rate due to temperature, [h-1]
KSX = 5 # Glucose saturation constant for the specific growth rate, [g L-1]
KOX = 0.0005 # Oxygen saturation constant for the specific growth rate, [g L-1]
qOmax = 0.05 # Maximum specific oxygen consumption rate, [h-1]
#Metabolic parameters
YPS = 0.51 # Theoretical yield of ethanol on glucose, [gP gS-1]
YXO = 0.97 # Theoretical yield of biomass on oxygen, [gX gO-1]
YXS = 0.53 # Theoretical yield of biomass on glucose, [gX gS-1]
#Physicochemical and thermodynamic parameters
Chbr = 4.18 # Heat capacity of the mass of reaction, [J g-1 oC-1]
Chc = 4.18 # Heat capacity of the cooling agent, [J g-1 oC-1]
DeltaH = 518000 # Heat of reaction of fermentation, [J mol-1 O2]
Tref = 20 # Reference temperature, [oC]
KH = 200 # Henry's constant for oxygen in the fermentation broth, [atm L mol-1]
z = 0.792 # Oxygen compressibility factor, [-]
R = 0.082 # Ideas gas constant, [L atm mol-1 oC-1]
kla0 = 100 # Temperature-independent volumetric oxygen transfer coefficient, [h-1]
KT = 360000 # Heat transfer coefficient, [J h-1 m-2 ??C-1]
rho = 1080 # Density of the fermentation broth, [g L-1]
rhoc = 1000 # Density of the cooling agent, [g L-1]
MO = 32.0 # Molecular weight of oxygen (O2), [g mol-1]
#Bioreactor design data
AT = 1.0 # Bioreactor heat transfer area, [m2]
V = 1800 # Bioreactor working volume, [L]
Vcj = 50 # Cooling jacket volume, [L]
Ogasin = 0.305 # Oxygen concentration in airflow inlet, [g L-1]
#Definition of model variables
#State variables
Xt = x[0] # Total cellular biomass, [g L-1]
Xv = x[1] # Viable cellular biomass, [g L-1]
S = x[2] # Substrate/Glucose concentration, [g L-1]
P = x[3] # Product/Ethanol concentration, [g L-1]
Oliq = x[4] # Dissolved oxygen concentration, [g L-1]
Ogas = x[5] # Gas phase oxygen (bubbles) in the fermentation broth, [g L-1]
T = x[6] # Temperature in the bioreactor, [oC]
Tc = x[7] # Temperature of the cooling agent in the jacket, [oC]
Vl = x[8] # Culture volume in the bioreactor, [L]
Sf_cum = x[9] # Cumulative amount of substrate/glucose fed to the bioreactor, [g]
Time = x[10] # Batch time, [h]
# Definition of model equations
# Kinetic rates
# -----------------------------
# Specific growth rate, [h-1]
mmax = ((a1*(T-k1))*(1-np.exp(b1*(T-k2))))**2
Pmax = Pmaxb + PmaxT/(1-np.exp(-b2*(T-k3)))
m1 = mmax * S/(KSX + S) * Oliq/(KOX + Oliq) * (1 - P/Pmax) * 1/(1+np.exp(-(100-S)/1)) # Specific growth rate, [h-1]
if m1 >= 0:
m = m1
else:
m=0.0
# Non-growth-associated ethanol specific production rate, [h-1]
if S > 0:
bP = c1 * np.exp(-AP1/T) - c2 * np.exp(-AP2/T) # Non-growth-associated ethanol specific production rate, [h-1]
else:
bP = 0.0
qP = aP*m + bP
# Ethanol consumption specific rate
qS = m/YXS + qP/YPS
# Oxygen consumption specific rate
qO = qOmax*Oliq/YXO/(KOX + Oliq)
# Specific biological deactivation rate of cell mass
Kd = Kdb + KdT/(1+np.exp(-b3*(T-k4)))
# Saturation concentration of oxygen in culture media
Osat = z*Ogas*R*T/KH
# Oxygen mass transfer coefficient
kla = kla0*1.2**(T-20)
# Volume of the gas phase in the bioreactor
Vg = V - Vl
#Material balances
#-----------------
# Volume of liquid culture
dVl = Qin - Qe
# Total cell mass
dXt = m*Xv + Qin/Vl*(Xtin-Xt)
# Total mass of biologically active cells
dXv = (m-Kd)*Xv + Qin/Vl*(Xvin-Xv)
# Glucose concentration
dS = Qin/Vl*(Sin-S) - qS*Xv
# Ethanol concentration
dP = Qin/Vl*(-P) + qP*Xv
# Disolved oxygen
dOliq = Qin/Vl*(Osat - Oliq) + kla*(Osat-Oliq) - qO*Xv
# Oxygen gas phase
dOgas = Fair/Vg*(Ogasin-Ogas) - Vl*kla/Vg*(Osat - Oliq) + Ogas*(Qin-Qe)/Vg
# Energy balances
#---------------
# Bioreactor temprature
dT = Qin/Vl*(Tin-T) - Tref/Vl*(Qin-Qe) + qO*Xv*DeltaH/MO/rho/Chbr - KT*AT*(T-Tc)/Vl/rho/Chbr
# Cooling agent temperature
dTc = Fc/Vcj*(Tcin-Tc) + KT*AT*(T-Tc)/Vcj/rhoc/Chc
# Yields & Productivity
#---------------------
# Cumulative amount of glucose fed to the bioreactor
dSf_cum = Sin*Qin
dTime = 1
# Definition of state derivatives vector
# State derivatives
dxdt = [dXt,dXv,dS,dP,dOliq,dOgas,dT,dTc,dVl,dSf_cum,dTime]
# [dxdt,mmax,Pmax,bP,m,Kd,Qin]
return dxdt
# test function
print(ethanol(x0,0.0,*ux0))
# Simulate the bioreactor operation until the selected time tf
x = odeint(ethanol,x0,tspan,args=ux0)
#plots Results
#Total and Viable Cellular Biomass
plt.figure()
plt.plot(tspan,x[:,0])
plt.plot(tspan,x[:,1])
plt.title('Total & Viable Cellular Biomass')
plt.ylabel('Biomass concentration [g/L]')
plt.xlabel('t [h]')
plt.legend(['Xt','Xv'])
plt.figure()
plt.title('Substrate (S) & Product (P) concentration')
plt.plot(tspan,x[:,2], label='S')
plt.plot(tspan,x[:,3], label='P')
plt.legend(); plt.grid()
plt.ylabel('Conc [g/L]')
plt.xlabel('Time [h]')
plt.minorticks_on()
plt.ylim(0)
plt.xlim(t[0],t[-1])
plt.tight_layout()
plt.figure()
plt.title('Bioreactor & Cooling jacket temperature')
plt.plot(tspan,x[:,6], label='T')
plt.plot(tspan,x[:,7], label='Tc')
plt.legend()
plt.ylabel('Temperature [oC]')
plt.xlabel('Time [h]')
plt.grid()
plt.minorticks_on()
plt.ylim(0)
plt.xlim(t[0],t[-1])
plt.tight_layout()
fig4, ax = plt.subplots()
ax.title.set_text('Dissolved & Gaseous Oxygen concentration')
lns1 = ax.plot(t,x[:,4], label='[Oliq]', color='c')
ax.set_xlabel('Time [h]')
ax.set_ylabel('Oliq [g/L]', color='c')
ax.minorticks_on()
ax2 = ax.twinx()
lns2 = ax2.plot(t,x[:,5], label='[Ogas]', color='y')
ax2.set_ylabel('Ogas [g/L]', color='y')
ax2.minorticks_on()
lns = lns1 + lns2
labs = [l.get_label() for l in lns]
ax.legend(lns, labs, loc='best')
ax.grid()
fig4.tight_layout()
plt.figure()
plt.title('Feeding Policy')
plt.plot(tspan, Qin, label='Qin')
plt.legend()
plt.ylabel('Qin [L/h]')
plt.xlabel('Time [h]')
plt.grid()
plt.minorticks_on()
plt.ylim(0)
plt.xlim(tspan[0],tspan[-1])
plt.tight_layout()
plt.show()页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/74362585
复制相关文章
相似问题

