如何将4个单独的数据集包装起来?
如何将4个单独的数据集包装起来?
提问于 2022-10-31 16:19:34
因此,我在尝试将4种不同的数据集包装起来时遇到了一些麻烦。每个数据集代表一个参与者,但我不知道如何组合这些数据集,这样就可以对它们进行面封装并共享x轴。以下是每个参与者的数据集:
p03
structure(list(day = structure(c(18880, 18880, 18908, 18908,
18911, 18912, 18912, 18913, 18913, 18914, 18914, 18915, 18915,
18916, 18916, 18917, 18917, 18918, 18918, 18919, 18920, 18921,
18921, 18922, 18922, 18923, 18924), class = "Date"), trialtype = c("go",
"nogo", "go", "nogo", "go", "go", "nogo", "go", "nogo", "go",
"nogo", "go", "nogo", "go", "nogo", "go", "nogo", "go", "nogo",
"go", "go", "go", "nogo", "go", "nogo", "go", "go"), mean = c(0.453363414634146,
0.21005, 0.63720350877193, 0.8809, 0.624077192982456, 0.581956842105263,
0.63065, 0.588196842105263, 0.607075, 0.540253289473684, 0.73215,
0.529153097345133, 0.583133333333333, 0.538794736842105, 0.4409,
0.522421428571429, 0.4792, 0.552129824561403, 0.5282, 0.512742105263158,
0.473664912280702, 0.528580701754386, 0.544266666666667, 0.490643859649123,
0.49955, 0.440738596491228, 0.475708771929825)), class = c("grouped_df",
"tbl_df", "tbl", "data.frame"), row.names = c(NA, -27L), groups = structure(list(
day = structure(c(18880, 18908, 18911, 18912, 18913, 18914,
18915, 18916, 18917, 18918, 18919, 18920, 18921, 18922, 18923,
18924), class = "Date"), .rows = structure(list(1:2, 3:4,
5L, 6:7, 8:9, 10:11, 12:13, 14:15, 16:17, 18:19, 20L,
21L, 22:23, 24:25, 26L, 27L), ptype = integer(0), class = c("vctrs_list_of",
"vctrs_vctr", "list"))), class = c("tbl_df", "tbl", "data.frame"
), row.names = c(NA, -16L), .drop = TRUE))p04
structure(list(day = structure(c(18880, 18880, 18918, 18918,
18919, 18920, 18922, 18923), class = "Date"), trialtype = c("go",
"nogo", "go", "nogo", "go", "go", "go", "go"), mean = c(0.453363414634146,
0.21005, 0.51259649122807, 0.5512, 0.631542105263158, 0.635184210526316,
0.718966666666667, 0.795638596491228)), class = c("grouped_df",
"tbl_df", "tbl", "data.frame"), row.names = c(NA, -8L), groups = structure(list(
day = structure(c(18880, 18918, 18919, 18920, 18922, 18923
), class = "Date"), .rows = structure(list(1:2, 3:4, 5L,
6L, 7L, 8L), ptype = integer(0), class = c("vctrs_list_of",
"vctrs_vctr", "list"))), class = c("tbl_df", "tbl", "data.frame"
), row.names = c(NA, -6L), .drop = TRUE))p05
structure(list(day = structure(c(18924, 18924, 18925, 18925,
18926, 18926, 18927, 18927, 18928, 18928, 18929, 18929, 18930,
18930, 18932, 18933, 18933, 18934, 18935, 18935, 18936, 18936,
18937, 18937, 18940, 18941, 18941, 18942, 18944, 18944, 18945,
18945, 18948, 18950, 18951, 18951), class = "Date"), trialtype = c("go",
"nogo", "go", "nogo", "go", "nogo", "go", "nogo", "go", "nogo",
"go", "nogo", "go", "nogo", "go", "go", "nogo", "go", "go", "nogo",
"go", "nogo", "go", "nogo", "go", "go", "nogo", "go", "go", "nogo",
"go", "nogo", "go", "go", "go", "nogo"), mean = c(0.693312280701754,
0.5485, 0.589122807017544, 1.26686666666667, 0.542644444444444,
0.3142, 0.607622807017544, 1.0739, 0.562494642857143, 0.45075,
0.522236842105263, 1.4405, 0.585782456140351, 0.574333333333333,
0.549415789473684, 0.499456140350877, 0.89835, 0.571619298245614,
0.836829824561404, 0.8192, 0.881107692307692, 2.16316666666667,
0.600214035087719, 0.6608, 0.665928571428571, 0.633136363636364,
0.4519, 0.558714035087719, 0.7840875, 1.20543333333333, 0.597352631578947,
1.564875, 0.517835087719298, 0.533582456140351, 0.526968421052632,
1.31243333333333)), class = c("grouped_df", "tbl_df", "tbl",
"data.frame"), row.names = c(NA, -36L), groups = structure(list(
day = structure(c(18924, 18925, 18926, 18927, 18928, 18929,
18930, 18932, 18933, 18934, 18935, 18936, 18937, 18940, 18941,
18942, 18944, 18945, 18948, 18950, 18951), class = "Date"),
.rows = structure(list(1:2, 3:4, 5:6, 7:8, 9:10, 11:12, 13:14,
15L, 16:17, 18L, 19:20, 21:22, 23:24, 25L, 26:27, 28L,
29:30, 31:32, 33L, 34L, 35:36), ptype = integer(0), class = c("vctrs_list_of",
"vctrs_vctr", "list"))), class = c("tbl_df", "tbl", "data.frame"
), row.names = c(NA, -21L), .drop = TRUE))p07
structure(list(day = structure(c(19210, 19210, 19211, 19211,
19212, 19212, 19213, 19213, 19214, 19214, 19215, 19215, 19216,
19216, 19217, 19217, 19218, 19218, 19219, 19221, 19222, 19223,
19223, 19224, 19225, 19226, 19226, 19227, 19227, 19228, 19228,
19229, 19229, 19230, 19231, 19231, 19232, 19232, 19238, 19238,
19239, 19239, 19240, 19241, 19242, 19242, 19243), class = "Date"),
trialtype = c("go", "nogo", "go", "nogo", "go", "nogo", "go",
"nogo", "go", "nogo", "go", "nogo", "go", "nogo", "go", "nogo",
"go", "nogo", "go", "go", "go", "go", "nogo", "go", "go",
"go", "nogo", "go", "nogo", "go", "nogo", "go", "nogo", "go",
"go", "nogo", "go", "nogo", "go", "nogo", "go", "nogo", "go",
"go", "go", "nogo", "go"), mean = c(0.496695906432749, 0.54765,
0.44894269005848, 0.406666666666667, 0.444126315789474, 0.3496,
0.442609356725146, 0.4528, 0.462447953216374, 0.4262, 0.47875730994152,
0.464266666666667, 0.459146783625731, 0.261666666666667,
0.479398830409357, 0.4199, 0.48975730994152, 0.2944, 0.511256140350877,
0.49047485380117, 0.543529239766082, 0.513159064327485, 0.5597,
0.51296783625731, 0.478275438596491, 0.492419298245614, 0.1061,
0.49356432748538, 0.4948, 0.534784795321637, 0.5097, 0.509980701754386,
0.2998, 0.505461988304094, 0.52029649122807, 0.52315, 0.52303216374269,
0.5337, 0.517447368421053, 0.3997, 0.517894152046784, 0.696366666666667,
0.511729824561403, 0.5346, 0.515273684210526, 0.508, 0.552337426900585
)), class = c("grouped_df", "tbl_df", "tbl", "data.frame"
), row.names = c(NA, -47L), groups = structure(list(day = structure(c(19210,
19211, 19212, 19213, 19214, 19215, 19216, 19217, 19218, 19219,
19221, 19222, 19223, 19224, 19225, 19226, 19227, 19228, 19229,
19230, 19231, 19232, 19238, 19239, 19240, 19241, 19242, 19243
), class = "Date"), .rows = structure(list(1:2, 3:4, 5:6, 7:8,
9:10, 11:12, 13:14, 15:16, 17:18, 19L, 20L, 21L, 22:23, 24L,
25L, 26:27, 28:29, 30:31, 32:33, 34L, 35:36, 37:38, 39:40,
41:42, 43L, 44L, 45:46, 47L), ptype = integer(0), class = c("vctrs_list_of",
"vctrs_vctr", "list"))), class = c("tbl_df", "tbl", "data.frame"
), row.names = c(NA, -28L), .drop = TRUE))从数据集中,我开始为每个参与者创建单独的数据可视化,如下所示:
p03_dropped_rt <- p03_response_times %>%
drop_na()
#convert day into date object
p03_dropped_rt$day <- as.Date(p03_dropped_rt$day)
head(p03_dropped_rt)
#create a line graph for p03 to show this data but have the x axis as just days
p3 <- p03_dropped_rt %>%
group_by(trialtype) %>%
arrange(day) %>%
mutate(days = paste0("day",row_number())) %>%
ggplot(aes(x = fct_inorder(days), y = mean, group = trialtype)) +
geom_line(aes(color = trialtype)) +
geom_point(aes(color = trialtype)) +
theme(axis.text.x = element_text(angle = 60, vjust = 0.5)) +
labs(title=" P03s Average response time of Go/No-go Trials",
x = "Day of training",
y = "Average Response Time (s) ",
color = "Trial Type")
p3
# repeat for p04
#drop missing values
p04_dropped_rt <- p04_response_times %>%
drop_na()
#convert day into date object
p04_dropped_rt$day <- as.Date(p04_dropped_rt$day)
head(p04_dropped_rt)
#create a line graph to show these results
p4 <- p04_dropped_rt %>%
group_by(trialtype) %>%
arrange(day) %>%
mutate(days = paste0("day",row_number())) %>%
ggplot(aes(x = fct_inorder(days), y = mean, group = trialtype)) +
geom_line(aes(color = trialtype)) +
geom_point(aes(color = trialtype)) +
theme(axis.text.x = element_text(angle = 60, vjust = 0.5)) +
labs(title=" P04s Average response time of Go/No-go Trials",
x = "Day of training",
y = "Average Response Time (s) ",
color = "Trial Type")
p4
#repeat for p05 and drop missing values
p05_dropped_rt <- p05_response_times %>%
drop_na()
#convert day into date object
p05_dropped_rt$day <- as.Date(p05_dropped_rt$day)
head(p05_dropped_rt)
#create a line graph to show this
p5 <- p05_dropped_rt %>%
group_by(trialtype) %>%
arrange(day) %>%
mutate(days = paste0("day",row_number())) %>%
ggplot(aes(x = fct_inorder(days), y = mean, group = trialtype)) +
geom_line(aes(color = trialtype)) +
geom_point(aes(color = trialtype)) +
theme(axis.text.x = element_text(angle = 60, vjust = 0.5)) +
labs(title=" P05s Average response time of Go/No-go Trials",
x = "Day of training",
y = "Average Response Time (s) ",
color = "Trial Type")
p5
# repeat for p07 and drop missing values
p07_dropped_rt <- p07_response_times %>%
drop_na()
# convert day to date object
p07_dropped_rt$day <- as.Date(p07_dropped_rt$day)
head(p07_dropped_rt)
#create a line graph to show this
p7 <- p07_dropped_rt %>%
group_by(trialtype) %>%
arrange(day) %>%
mutate(days = paste0("day",row_number())) %>%
ggplot(aes(x = fct_inorder(days), y = mean, group = trialtype)) +
geom_line(aes(color = trialtype)) +
geom_point(aes(color = trialtype)) +
theme(axis.text.x = element_text(angle = 60, vjust = 0.5)) +
labs(title=" P07s Average response time of Go/No-go Trials",
x = "Day of training",
y = "Average Response Time (s) ",
color = "Trial Type")
p7我不知道该如何包装,所以我尝试了‘gg排列’。然而,这导致图形被令人难以置信地展开,并没有外观良好的视觉效果。如果你想看的话,我在最后附了一张照片。
有人能帮我吗?
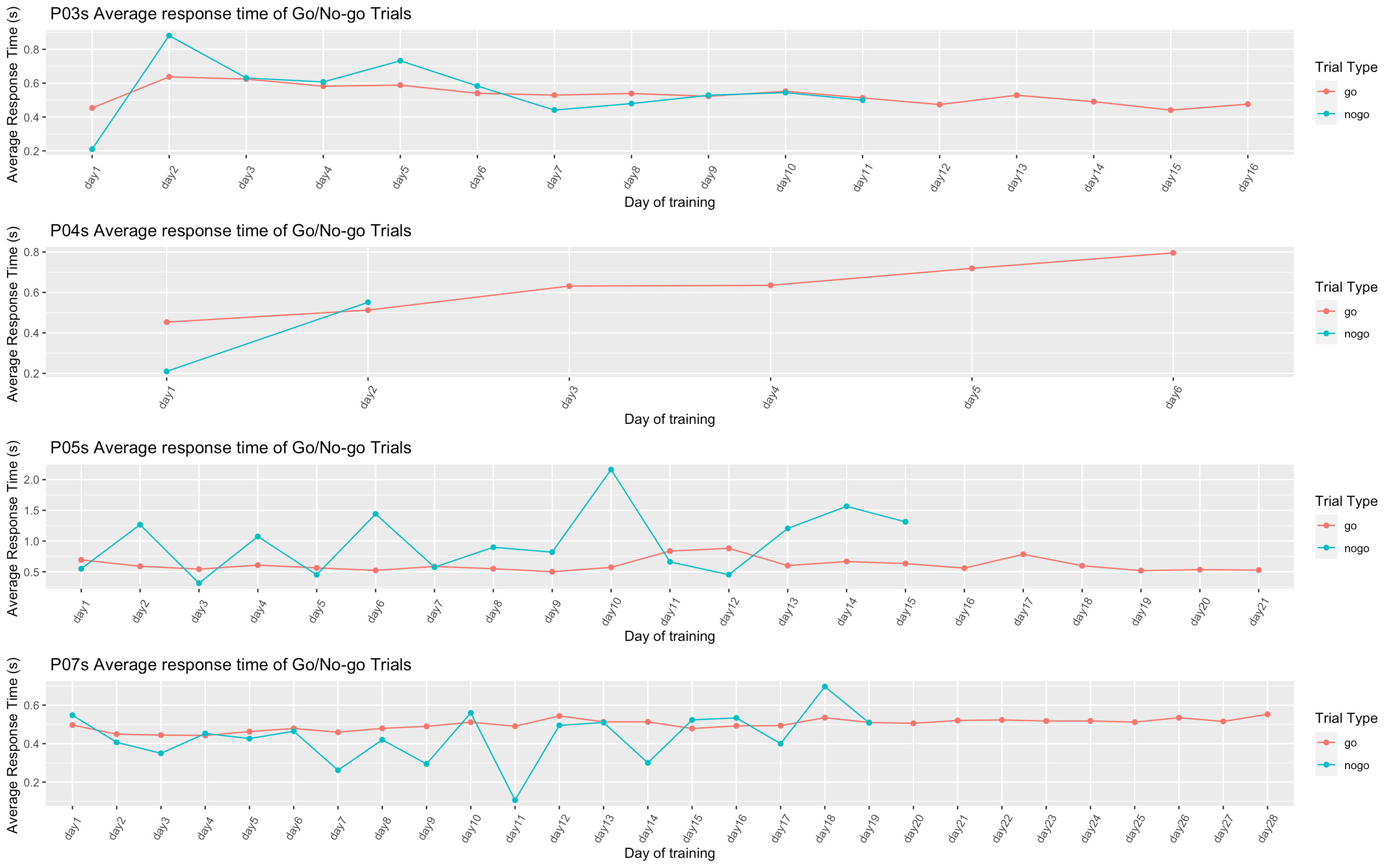
我试图按以下方式绑定这些行
bind_rows(list(p3, p4, p5, p7), .id="patient") %>%
group_by(trialtype) %>%
arrange(day) %>%
mutate(days = paste0("day",row_number())) %>%
ggplot(aes(x = fct_inorder(days), y = mean, group = trialtype)) +
geom_line(aes(color = trialtype)) +
geom_point(aes(color = trialtype)) +
theme(axis.text.x = element_text(angle = 60, vjust = 0.5)) +
labs(title="Average response time of Go/No-go Trials",
x = "Day of training",
y = "Average Response Time (s) ",
color = "Trial Type") + facet_wrap(patient)它产生了这个错误
Error in `bind_rows()`:
! Argument 1 must be a data frame or a named atomic vector.回答 1
Stack Overflow用户
发布于 2022-10-31 21:24:48
我不知道你在做什么,但显然是不对的。
bind_rows(list(p03, p04, p05, p07), .id="patient") %>%
group_by(trialtype) %>%
arrange(day) %>%
mutate(days = paste0("day",row_number())) %>%
ggplot(aes(x = fct_inorder(days), y = mean, group = trialtype)) +
geom_line(aes(color = trialtype)) +
geom_point(aes(color = trialtype)) +
theme(axis.text.x = element_text(angle = 60, vjust = 0.5)) +
labs(title="Average response time of Go/No-go Trials",
x = "Day of training",
y = "Average Response Time (s) ",
color = "Trial Type") +
facet_wrap(vars(patient))给出
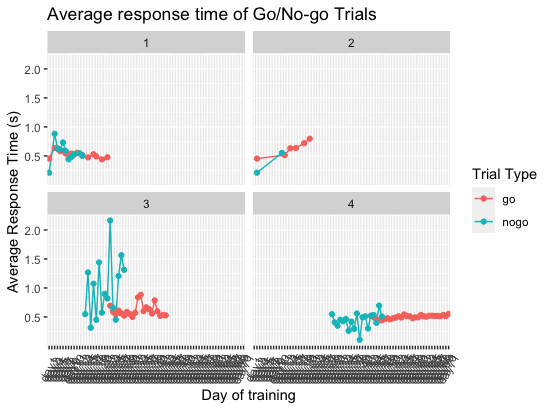
若要访问联机帮助,请使用?。例如,> ?facet_wrap。或者在RStudio帮助面板顶部的搜索栏中键入函数的名称。或者使用谷歌。
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/74266228
复制相关文章
相似问题

