动力系统分岔图
动力系统分岔图
提问于 2022-10-09 20:28:28
TL:博士
如何在Python中实现季节性强制流行病学模型的分叉图,例如SEIR (易感、暴露、感染、恢复)?我已经知道如何实现模型本身并显示抽样时间序列(参见这个堆叠溢出问题),但我很难从教科书中复制分叉图。
上下文与我的尝试
我试图复制“人类和动物传染病建模”(Keeling,2007)一书中的数字,以验证我对模型的实现,并学习/可视化不同的模型参数如何影响动力系统的演化。以下是教科书上的数字。
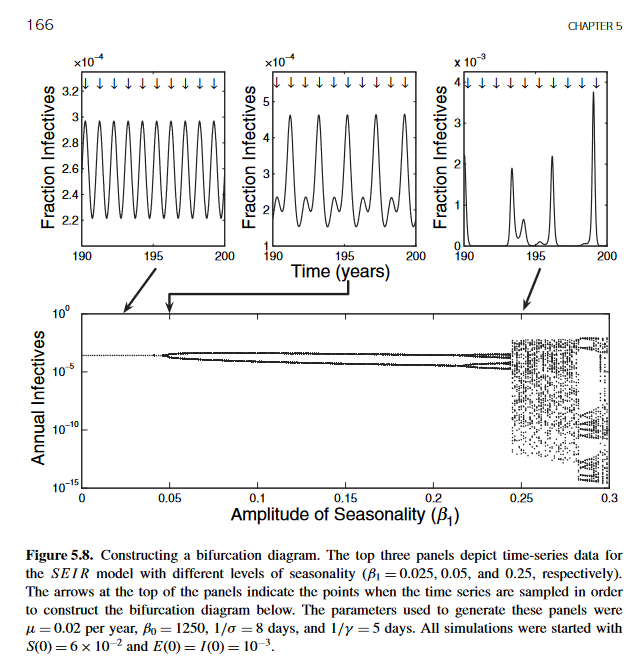
我已经为使用逻辑映射的例子找到了分支图的实现(请参阅这个ipython食谱 - 肾盂分叉,和这个堆栈过流问题)。我从这些实现中得到的主要结论是,分岔图上的一个点具有一个x分量,它的x分量等于可变参数的某个特定值(例如,Beta 1= 0.025),并且它的y分量是给定模型/函数在时间t处的解(数值或其他)。在这个问题的末尾,我使用这个逻辑实现代码部分中的plot_bifurcation函数。
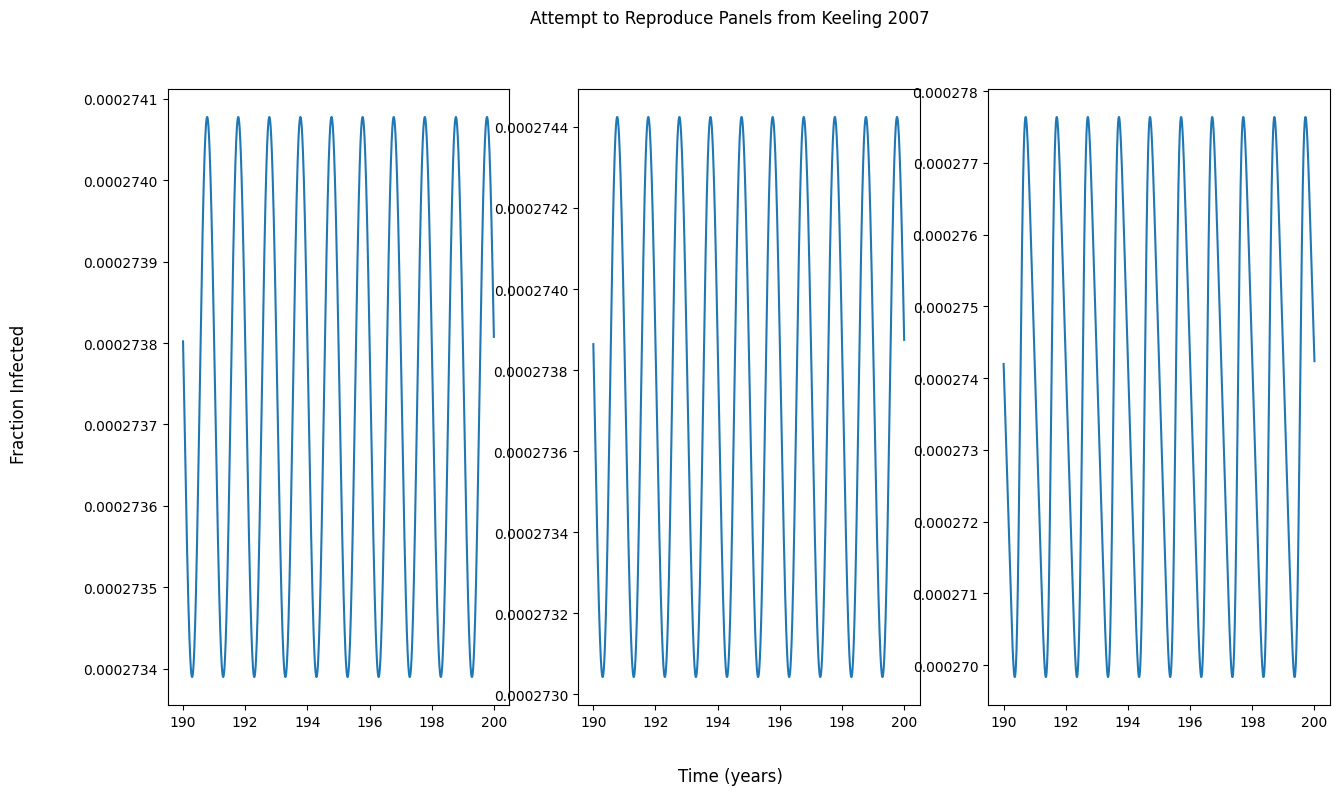
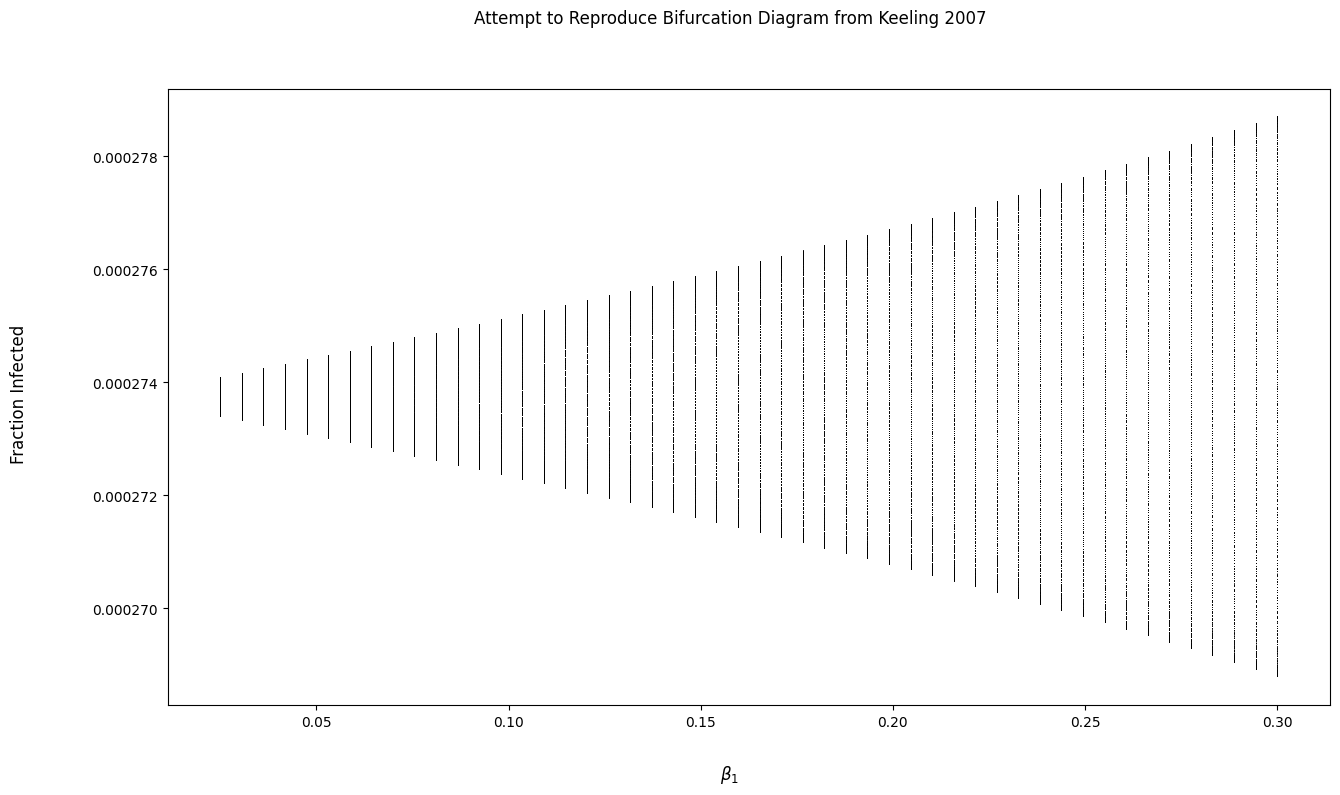
问题
为什么我的面板输出与图中的不匹配?我假设,如果没有面板匹配教科书中的输出,我就无法从教科书中复制分叉图。
我试图实现一个函数来生成分岔图,但是输出看起来真的很奇怪。我对分叉图有什么误解吗?
注意:在代码执行期间,我没有收到警告/错误。
代码复制我的数字
from typing import Callable, Dict, List, Optional, Any
import numpy as np
import matplotlib.pyplot as plt
from scipy.integrate import odeint
def seasonal_seir(y: List, t: List, params: Dict[str, Any]):
"""Seasonally forced SEIR model.
Function parameters much match with those required
by `scipy.integrate.odeint`
Args:
y: Initial conditions.
t: Timesteps over which numerical solution will be computed.
params: Dict with the following key-value pairs:
beta_zero -- Average transmission rate.
beta_one -- Amplitude of seasonal forcing.
omega -- Period of forcing.
mu -- Natural mortality rate.
sigma -- Latent period for infection.
gamma -- Recovery from infection term.
Returns:
Tuple whose components are the derivatives of the
susceptible, exposed, and infected state variables
w.r.t to time.
References:
[SEIR Python Program from Textbook](http://homepages.warwick.ac.uk/~masfz/ModelingInfectiousDiseases/Chapter2/Program_2.6/Program_2_6.py)
[Seasonally Forced SIR Program from Textbook](http://homepages.warwick.ac.uk/~masfz/ModelingInfectiousDiseases/Chapter5/Program_5.1/Program_5_1.py)
"""
beta_zero = params['beta_zero']
beta_one = params['beta_one']
omega = params['omega']
mu = params['mu']
sigma = params['sigma']
gamma = params['gamma']
s, e, i = y
beta = beta_zero*(1 + beta_one*np.cos(omega*t))
sdot = mu - (beta * i + mu)*s
edot = beta*s*i - (mu + sigma)*e
idot = sigma*e - (mu + gamma)*i
return sdot, edot, idot
def plot_panels(
model: Callable,
model_params: Dict,
panel_param_space: List,
panel_param_name: str,
initial_conditions: List,
timesteps: List,
odeint_kwargs: Optional[Dict] = dict(),
x_ticks: Optional[List] = None,
time_slice: Optional[slice] = None,
state_var_ix: Optional[int] = None,
log_scale: bool = False):
"""Plot panels that are samples of the parameter space for bifurcation.
Args:
model: Function that models dynamical system. Returns dydt.
model_params: Dict whose key-value pairs are the names
of parameters in a given model and the values of those parameters.
bifurcation_parameter_space: List of varied bifurcation parameters.
bifuraction_parameter_name: The name o the bifurcation parameter.
initial_conditions: Initial conditions for numerical integration.
timesteps: Timesteps for numerical integration.
odeint_kwargs: Key word args for numerical integration.
state_var_ix: State variable in solutions to use for plot.
time_slice: Restrict the bifurcation plot to a subset
of the all solutions for numerical integration timestep space.
Returns:
Figure and axes tuple.
"""
# Set default ticks
if x_ticks is None:
x_ticks = timesteps
# Create figure
fig, axs = plt.subplots(ncols=len(panel_param_space))
# For each parameter that is varied for a given panel
# compute numerical solutions and plot
for ix, panel_param in enumerate(panel_param_space):
# update model parameters with the varied parameter
model_params[panel_param_name] = panel_param
# Compute solutions
solutions = odeint(
model,
initial_conditions,
timesteps,
args=(model_params,),
**odeint_kwargs)
# If there is a particular solution of interst, index it
# otherwise squeeze last dimension so that [T, 1] --> [T]
# where T is the max number of timesteps
if state_var_ix is not None:
solutions = solutions[:, state_var_ix]
elif state_var_ix is None and solutions.shape[-1] == 1:
solutions = np.squeeze(solutions)
else:
raise ValueError(
f'solutions to model are rank-2 tensor of shape {solutions.shape}'
' with the second dimension greater than 1. You must pass'
' a value to :param state_var_ix:')
# Slice the solutions based on the desired time range
if time_slice is not None:
solutions = solutions[time_slice]
# Natural log scale the results
if log_scale:
solutions = np.log(solutions)
# Plot the results
axs[ix].plot(x_ticks, solutions)
return fig, axs
def plot_bifurcation(
model: Callable,
model_params: Dict,
bifurcation_parameter_space: List,
bifurcation_param_name: str,
initial_conditions: List,
timesteps: List,
odeint_kwargs: Optional[Dict] = dict(),
state_var_ix: Optional[int] = None,
time_slice: Optional[slice] = None,
log_scale: bool = False):
"""Plot a bifurcation diagram of state variable from dynamical system.
Args:
model: Function that models system. Returns dydt.
model_params: Dict whose key-value pairs are the names
of parameters in a given model and the values of those parameters.
bifurcation_parameter_space: List of varied bifurcation parameters.
bifuraction_parameter_name: The name o the bifurcation parameter.
initial_conditions: Initial conditions for numerical integration.
timesteps: Timesteps for numerical integration.
odeint_kwargs: Key word args for numerical integration.
state_var_ix: State variable in solutions to use for plot.
time_slice: Restrict the bifurcation plot to a subset
of the all solutions for numerical integration timestep space.
log_scale: Flag to natural log scale solutions.
Returns:
Figure and axes tuple.
"""
# Track the solutions for each parameter
parameter_x_time_matrix = []
# Iterate through parameters
for param in bifurcation_parameter_space:
# Update the parameter dictionary for the model
model_params[bifurcation_param_name] = param
# Compute the solutions to the model using
# dictionary of parameters (including the bifurcation parameter)
solutions = odeint(
model,
initial_conditions,
timesteps,
args=(model_params, ),
**odeint_kwargs)
# If there is a particular solution of interst, index it
# otherwise squeeze last dimension so that [T, 1] --> [T]
# where T is the max number of timesteps
if state_var_ix is not None:
solutions = solutions[:, state_var_ix]
elif state_var_ix is None and solutions.shape[-1] == 1:
solutions = np.squeeze(solutions)
else:
raise ValueError(
f'solutions to model are rank-2 tensor of shape {solutions.shape}'
' with the second dimension greater than 1. You must pass'
' a value to :param state_var_ix:')
# Update the parent list of solutions for this particular
# bifurcation parameter
parameter_x_time_matrix.append(solutions)
# Cast to numpy array
parameter_x_time_matrix = np.array(parameter_x_time_matrix)
# Transpose: Bifurcation plots Function Output vs. Parameter
# This line ensures that each row in the matrix is the solution
# to a particular state variable in the system of ODEs
# a timestep t
# and each column is that solution for a particular value of
# the (varied) bifurcation parameter of interest
time_x_parameter_matrix = np.transpose(parameter_x_time_matrix)
# Slice the iterations to display to a smaller range
if time_slice is not None:
time_x_parameter_matrix = time_x_parameter_matrix[time_slice]
# Make bifurcation plot
fig, ax = plt.subplots()
# For the solutions vector at timestep plot the bifurcation
# NOTE: The elements of the solutions vector represent the
# numerical solutions at timestep t for all varied parameters
# in the parameter space
# e.g.,
# t beta1=0.025 beta1=0.030 .... beta1=0.30
# 0 solution00 solution01 .... solution0P
for sol_at_time_t_for_all_params in time_x_parameter_matrix:
if log_scale:
sol_at_time_t_for_all_params = np.log(sol_at_time_t_for_all_params)
ax.plot(
bifurcation_parameter_space,
sol_at_time_t_for_all_params,
',k',
alpha=0.25)
return fig, ax
# Define initial conditions based on figure
s0 = 6e-2
e0 = i0 = 1e-3
initial_conditions = [s0, e0, i0]
# Define model parameters based on figure
# NOTE: omega is not mentioned in the figure, but
# omega is defined elsewhere as 2pi/365
days_per_year = 365
mu = 0.02/days_per_year
beta_zero = 1250
sigma = 1/8
gamma = 1/5
omega = 2*np.pi / days_per_year
model_params = dict(
beta_zero=beta_zero,
omega=omega,
mu=mu,
sigma=sigma,
gamma=gamma)
# Define timesteps
nyears = 200
ndays = nyears * days_per_year
timesteps = np.arange(1, ndays + 1, 1)
# Define different levels of seasonality (from figure)
beta_ones = [0.025, 0.05, 0.25]
# Define the time range to actually show on the plot
min_year = 190
max_year = 200
# Create a slice of the iterations to display on the diagram
time_slice = slice(min_year*days_per_year, max_year*days_per_year)
# Get the xticks to display on the plot based on the time slice
x_ticks = timesteps[time_slice]/days_per_year
# Plot the panels using the infected state variable ix
infection_ix = 2
# Plot the panels
panel_fig, panel_ax = plot_panels(
model=seasonal_seir,
model_params=model_params,
panel_param_space=beta_ones,
panel_param_name='beta_one',
initial_conditions=initial_conditions,
timesteps=timesteps,
odeint_kwargs=dict(hmax=5),
x_ticks=x_ticks,
time_slice=time_slice,
state_var_ix=infection_ix,
log_scale=False)
# Label the panels
panel_fig.suptitle('Attempt to Reproduce Panels from Keeling 2007')
panel_fig.supxlabel('Time (years)')
panel_fig.supylabel('Fraction Infected')
panel_fig.set_size_inches(15, 8)
# Plot bifurcation
bi_fig, bi_ax = plot_bifurcation(
model=seasonal_seir,
model_params=model_params,
bifurcation_parameter_space=np.linspace(0.025, 0.3),
bifurcation_param_name='beta_one',
initial_conditions=initial_conditions,
timesteps=timesteps,
odeint_kwargs={'hmax':5},
state_var_ix=infection_ix,
time_slice=time_slice,
log_scale=False)
# Label the bifurcation
bi_fig.suptitle('Attempt to Reproduce Bifurcation Diagram from Keeling 2007')
bi_fig.supxlabel(r'$\beta_1$')
bi_fig.supylabel('Fraction Infected')
bi_fig.set_size_inches(15, 8)回答 1
Stack Overflow用户
回答已采纳
发布于 2022-10-11 16:28:56
这个问题的答案是计算科学堆栈交换上的这里。都归功于卢茨·莱曼。
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/74008146
复制相关文章
相似问题

