图形化数据的变化顺序
图形化数据的变化顺序
提问于 2022-08-03 13:48:05
我试图按照表中指定的顺序绘制我的数据,但我不知道如何绘制。我想我已经改变了水平(?)但不是因素(?)
链接到我的数据:https://docs.google.com/spreadsheets/d/1juBhHNjvsm1otxcpEFu-Zi_v5hSrNj94zdXrENOglTs/edit?usp=sharing
到目前为止,我已经:
BigTree <- read_csv(file = "/cloud/project/Phylogeny/RtreeData.csv")
Bfr <- BigTree[with(BigTree, order(-BfrA, -BfrB, -Bfd, -Ftn, -DPS,
-Rubrerythrin, -EncFtn, -NEEF)), ]
BfrLong <- pivot_longer(Bfr, 2:9)
colnames(BfrLong) <- c("Species", "Protein", "Presence")这可以重新排序数据,并按照我想要绘制的顺序放置数据,但是我无法绘制这个数据。错误说
错误(BfrLong,aes(x =蛋白质,y=物种,fill =存在))+:二进制运算符的非数值参数
我试着使用下面的因子函数得到相同的结果,但是它没有达到预期的效果。我在这里做错什么了?
BigTreeLong$Protein <- factor(BigTreeLong$Protein,
levels = c("BfrA", "BfrB", "Bfd", "Ftn", "DPS",
"Rubrerythrin", "EncFtn", "NEEF"))如何更改因素,以便在绘图期间维护顺序?
到目前为止,我的热图代码使用了原始的BigTree数据格式。我想排序,所以左上角开始最浅绿色,它逐渐减少到大多数深绿色的底部行。
ggplot(BigTreeLong, aes(Protein, Species, fill= Presence)) +
geom_tile(color = "white", linetype = 1) +
scale_fill_gradient(low = "darkgreen", high = "olivedrab3") +
labs(title = "Ferritins present in different bacteria",
x = "Ferritin family proteins",
y = "Species",
fill = "Presence of protein") +
theme_minimal() +
theme(axis.text.x = element_text(angle=45, hjust=1, size=8)) +
theme(axis.text.y = element_text(angle=0, hjust=1, size=6)) +
scale_x_discrete(limits = c("BfrA", "BfrB", "Bfd", "Ftn", "DPS",
"EncFtn", "NEEF", "Rubrerythrin"))这是我目前制作的热图
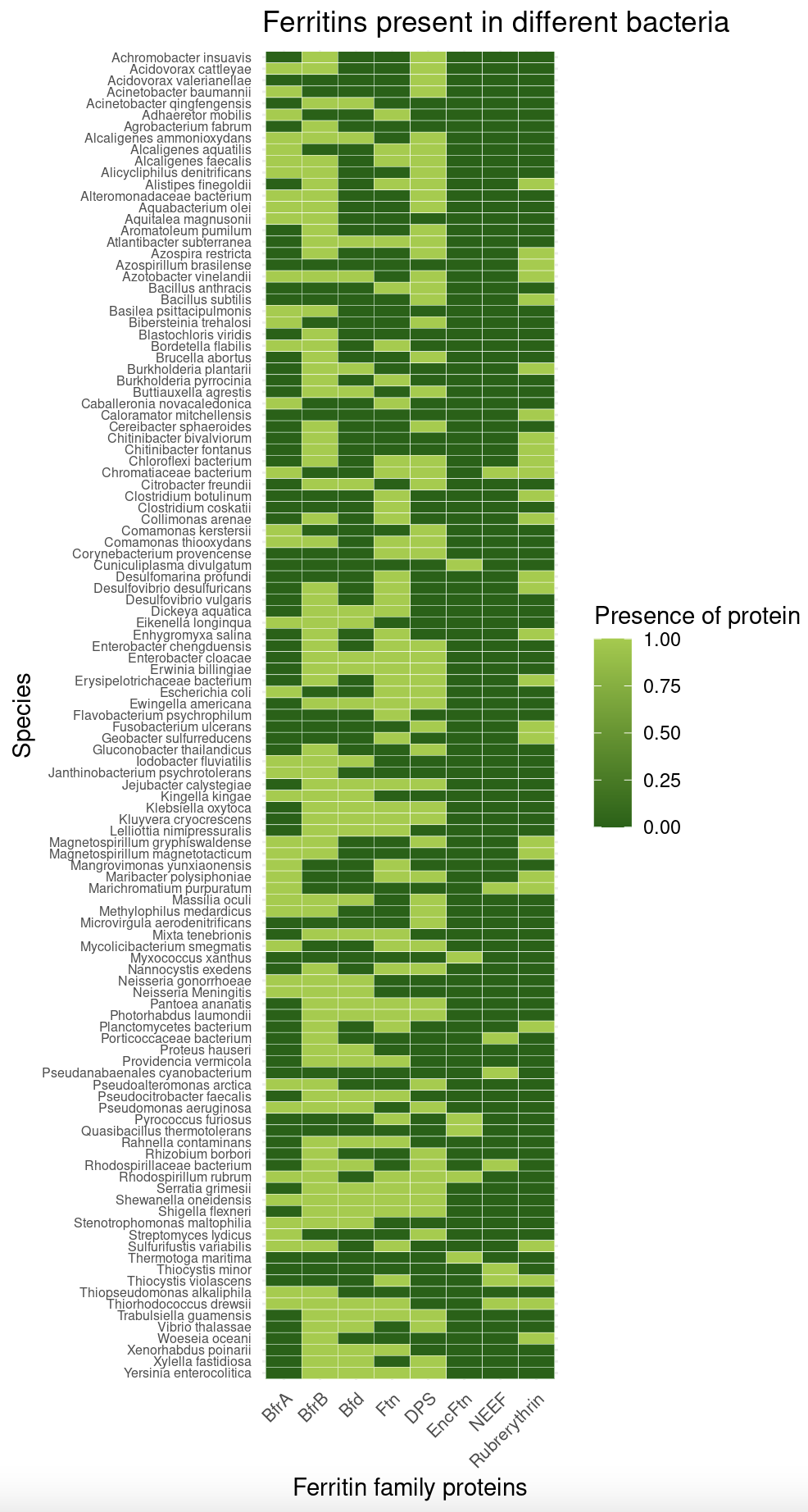
我希望我已经包括了我需要的一切!
回答 1
Stack Overflow用户
回答已采纳
发布于 2022-08-03 14:05:22
您的物种列的级别在您的代码中没有被修改,所以您需要这一行来注册它:
BfrLong$Species=factor(BfrLong$Species,levels=Bfr$Species)请注意示例中对象的名称,有BfrLong和BigTreeLong,但这应该有效:
BigTree <- read_csv(file = "/cloud/project/Phylogeny/RtreeData.csv")
Bfr <- BigTree[with(BigTree, order(-BfrA, -BfrB, -Bfd, -Ftn, -DPS,
-Rubrerythrin, -EncFtn, -NEEF)), ]
BfrLong <- pivot_longer(Bfr, 2:9)
colnames(BfrLong) <- c("Species", "Protein", "Presence")
BfrLong$Species=factor(BfrLong$Species,levels=Bfr$Species)
ggplot(BfrLong, aes(Protein, Species, fill= Presence)) +
geom_tile(color = "white", linetype = 1) +
scale_fill_gradient(low = "darkgreen", high = "olivedrab3") +
labs(title = "Ferritins present in different bacteria",
x = "Ferritin family proteins",
y = "Species",
fill = "Presence of protein") +
theme_minimal() +
theme(axis.text.x = element_text(angle=45, hjust=1, size=8)) +
theme(axis.text.y = element_text(angle=0, hjust=1, size=6)) +
scale_x_discrete(limits = c("BfrA", "BfrB", "Bfd", "Ftn", "DPS",
"EncFtn", "NEEF", "Rubrerythrin"))
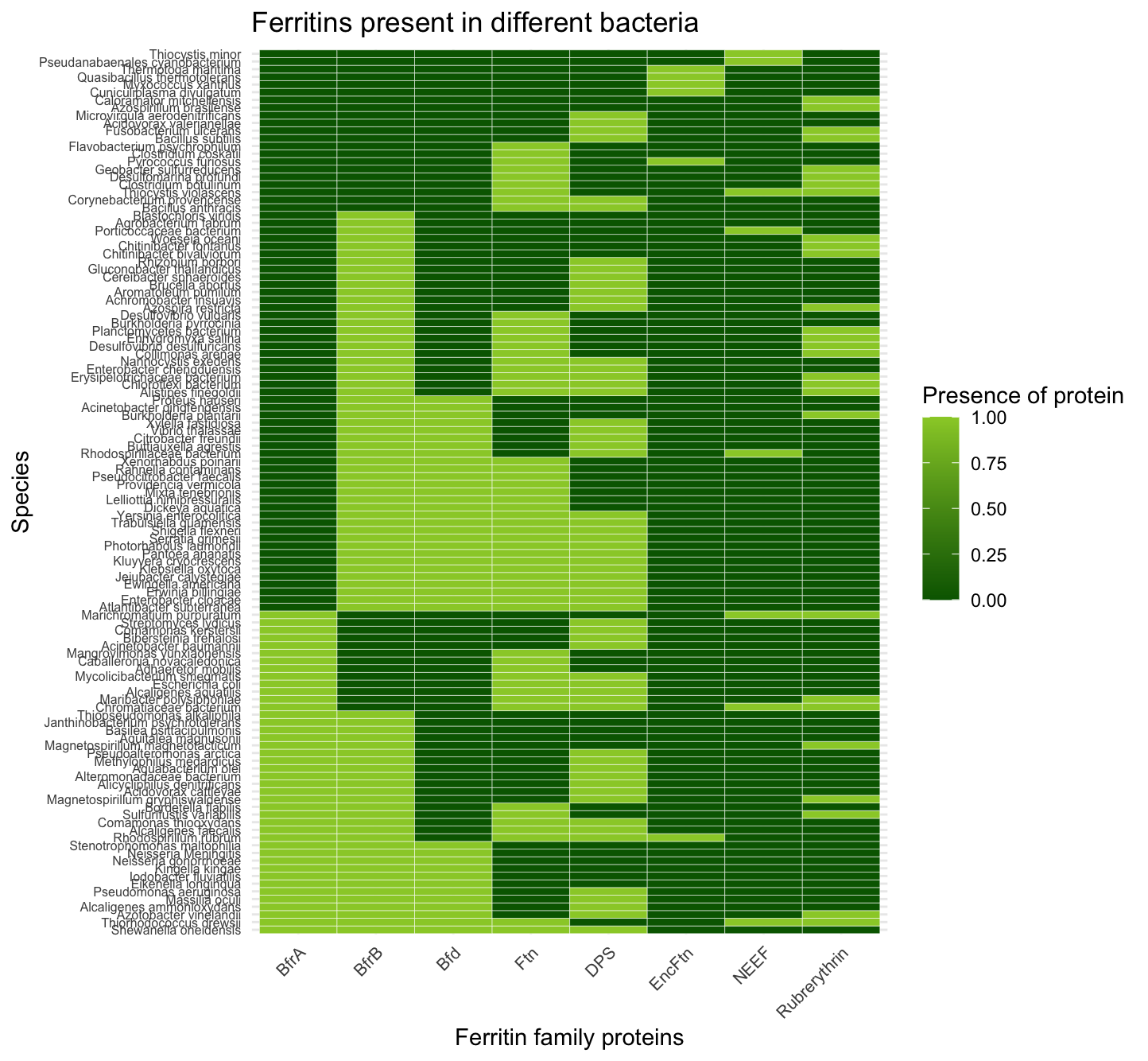
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/73222808
复制相关文章
相似问题

