R中3个PDE的细胞生长模型(Monod)拟合
R中3个PDE的细胞生长模型(Monod)拟合
提问于 2022-07-29 17:15:31
用@tpetzoldt的答案解决了问题,对原来的代码进行了修改以成功地运行fit。
大家好,我试着用3个偏微分方程来拟合实验数据,找出4个系数,包括mumax,Ks,Y_(x/s)和Y_(p/s)。我使用的代码处理了2组PDE,但没有处理这组3。
需要安装的一组PDE:
dX/dt =木乃伊。S.X/ (Ks + S)
dS/dt = -1/Y_(x/s)木乃伊。S.X/ (Ks + S)
dP/dt =α。dX/dt +β。X
library(deSolve)
library(ggplot2)
library(minpack.lm)
library(reshape2)
time <- c(0, 3, 5, 8, 9.5, 11.5, 14, 16, 18, 20, 25, 27)
X <- c(0.0904, 0.1503, 0.2407, 0.3864, 0.5201, 0.6667, 0.8159, 0.9979, 1.0673, 1.1224, 1.1512, 1.2093)
S <- c(9.0115, 8.8088, 7.9229, 7.2668, 5.3347, 4.911, 3.5354, 1.4041, 0, 0, 0, 0)
P <- c(0.0151, 0.0328, 0.0621, 0.1259, 0.2949, 0.3715, 0.4199, 0.522, 0.5345, 0.6081, 0.07662, 0.7869)
data <- data.frame(time, X, S, P)
Monod <- function(t,c,parms) {
k1 <- parms$k1 # mumax
k2 <- parms$k2 # Ks
k3 <- parms$k3 # Y_(X/S)
k4 <- parms$k4 # alpha
k5 <- parms$k5 # beta
r <- numeric(length(c))
r[1] <- k1 * c["S"] * c["X"] / ( k2 + c["S"] ) # r[1] = dCx/dt = k1.Cs.Cx/(k2+Cs)
r[2] <- -k1 * c["S"] * c["X"] / ( ( k2 + c["S"] ) * k3 ) # r[2] = dCs/dt = -1/k3 * dCx/dt
r[3] <- k4 * r[1] + k5 * c["X"] # r[3] = dCp/dt = alpha * dX/dt + beta * X
return(list(r))
}
residuals <- function(parms){
cinit <- c( X=data[1,2], S=data[1,3], P=data[1,4] )
t <- c(seq(0, 27, 1), data$time)
t <- sort(unique(t))
k1 <- parms[1]
k2 <- parms[2]
k3 <- parms[3]
k4 <- parms[4]
k5 <- parms[5]
out <- ode( y = cinit,
times = t,
func = Monod,
parms = list( k1 = k1, k2 = k2, k3 = k3, k4 = k4, k5 = k5) )
out_Monod <- data.frame(out)
out_Monod <- out_Monod[out_Monod$t %in% data$time,]
pred_Monod <- melt(out_Monod,id.var="time",variable.name="Substance",value.name="Conc")
exp_Monod <- melt(data,id.var="time",variable.name="Substance",value.name="Conc")
residuals <- pred_Monod$Conc-exp_Monod$Conc
return(residuals)
}
parms <- c(k1=0.5, k2=6.5, k3=0.2, k4=1.2, k5 = 0.1)
fitval <- nls.lm(par=parms,fn=residuals)
cinit <- c(X=data[1,2], S=data[1,3], P=data[1,4])
out <- ode(y=cinit, times=seq(min(data$time), max(data$time)), func = Monod, parms=as.list(coef(fitval)))
plot(out, obs=data, mfrow=c(1, 3))回答 1
Stack Overflow用户
回答已采纳
发布于 2022-07-31 10:01:40
原来的守则有几个问题:
output
- concentration中的状态变量在数据中为小写,在模型中为大小写,在初始值中为无任何名称,在数据中为
- 时间,在ode中为
time,在另一个地方为Concentration,而在另一个
H 111中,名称ssq和D13具有误导性,因为函数计算残差而不是平方和。在背景中计算平方和(参见?nls.lm)
the赋值运算符<-和=的使用不一致)。这不是一个错误,但是人工名称的style.
instead并不是很好--可以直接使用原始名称如mumax或alpha.
It --不清楚为什么要使用data2或Monod2这样的名称,因为没有data1.
。
解决了所有这些问题,我们得出了下面的解决方案。它通过和适合某种方式,但它仍然可以改进。这些警告目前可能被忽略。原因是,参数超出了允许的范围。当然,这是可以解决的,但这将是另一个问题。
library(deSolve)
library(ggplot2)
library(minpack.lm)
library(reshape2)
data <- data.frame(
time = c(0, 3, 5, 8, 9.5, 11.5, 14, 16, 18, 20, 25, 27),
X = c(0.0904, 0.1503, 0.2407, 0.3864, 0.5201, 0.6667, 0.8159, 0.9979, 1.0673, 1.1224, 1.1512, 1.2093),
S = c(9.0115, 8.8088, 7.9229, 7.2668, 5.3347, 4.911, 3.5354, 1.4041, 0, 0, 0, 0),
P = c(0.0151, 0.0328, 0.0621, 0.1259, 0.2949, 0.3715, 0.4199, 0.522, 0.5345, 0.6081, 0.07662, 0.7869)
)
Monod <- function(t, c, parms) {
k1 <- parms$k1 # mumax
k2 <- parms$k2 # Ks
k3 <- parms$k3 # Y_(X/S)
k4 <- parms$k4 # alpha
k5 <- parms$k4 # beta
r <- numeric(length(c))
r[1] <- k1 * c["S"] * c["X"] / (k2 + c["S"])
r[2] <- -k1 * c["S"] * c["X"] / ((k2 + c["S"]) * k3)
r[3] <- k4 * r[1] + k5 * c["X"]
return(list(r))
}
cinit <- c(X = data[1, 2], S = data[1, 3], P = data[1, 4])
t <- seq(0, 27, 1)
parms <- list(k1=0.1, k2=5, k3=1, k4=0.08, k5 = 0.005)
out <- ode(y=cinit, times=t, func = Monod, parms=parms)
plot(out)
residuals <- function(parms){
cinit <- c(X = data[1, 2], S = data[1, 3], P = data[1, 4] )
# time points for which conc is reported including the points where data is available
t <- c(seq(0, 27, 1), data$t)
t <- sort(unique(t))
# parameters from the parameter estimation routine:
k1 <- parms[1]
k2 <- parms[2]
k3 <- parms[3]
k4 <- parms[4]
# solve ODE for a given set of parameters
out <- ode(y = cinit, times = t, func = Monod, parms = list( k1 = k1, k2 = k2, k3 = k3, k4 = k4) )
# Filter data that contains time points where data is available:
out_Monod <- data.frame(out)
out_Monod <- out_Monod[out_Monod$t %in% data$t,]
# Evaluate predicted vs experimental residual
pred_Monod <- melt(out_Monod, id.var="time", variable.name="Substance", value.name="conc")
exp_Monod <- melt(data, id.var="time", variable.name="Substance", value.name="conc")
residuals <- pred_Monod$conc - exp_Monod$conc
# return predicted vs experimental residual
return(residuals)
}
# initial guess for parameters
parms <- c(k1=0.5, k2=6.5, k3=0.2, k4=1.2)
fitval <- nls.lm(par=parms, fn=residuals)
summary(fitval)
out <- ode(y=cinit, times=seq(min(data$time), max(data$time)), func = Monod, parms=as.list(coef(fitval)))
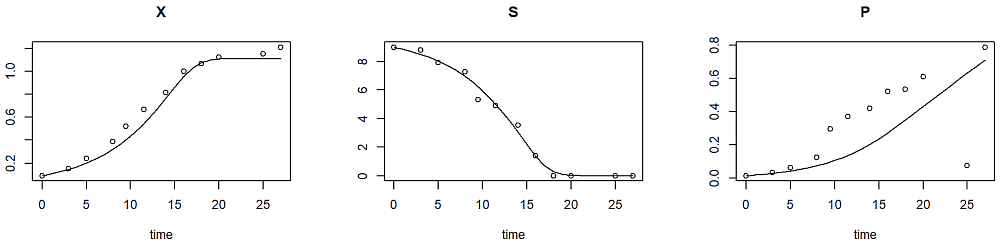
plot(out, obs=data, mfrow=c(1, 3))
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/73168906
复制相关文章
相似问题

