删除shapefile
删除shapefile
提问于 2022-05-05 04:05:17
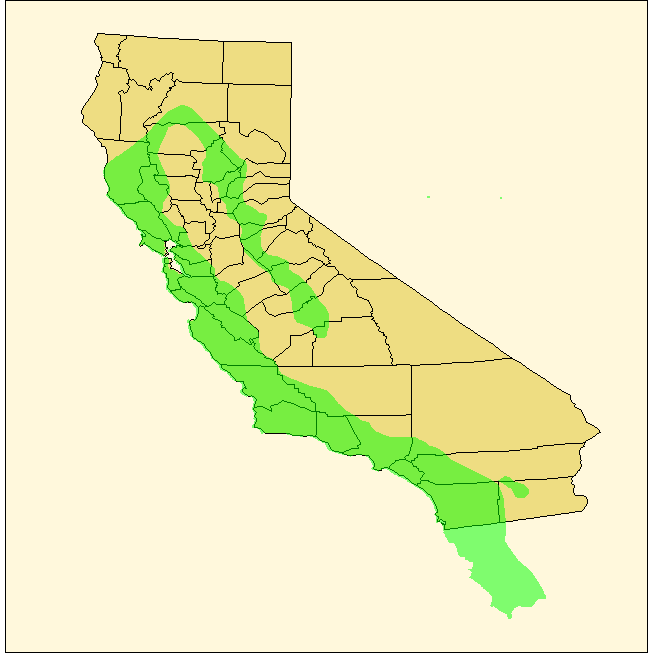
我在这里有一个形状文件:https://login.filesanywhere.com/fs/v.aspx?v=8c6c63865a6574bcaa69,我有一个加州红腿青蛙的形状文件,我正在加州的顶部,然而,这些青蛙的范围延伸到加州以外的地方,进入墨西哥。我只想要加州的青蛙数据,我怎样才能修剪扩展到墨西哥的数据呢?我试着使用子集来分隔“原产地”,但它似乎没有任何效果。谢谢你事先提供的帮助。
library(rgdal)
library(tidyverse)
ranas <- readOGR(dsn = ".", layer = "data_0")
names(ranas)
# Coerce into a dataframe
ranas4 <- fortify(ranas)
head(ranas4)
cali_map <- map_data("state",region="california")
counties <- map_data("county",region="California")
head(counties)
windows(w=9)
ggplot(cali_map, aes(x = long, y = lat, group = group)) +
geom_polygon() +
geom_polygon(data = ranas4, fill = "green")
ggplot(cali_map, aes(x = long, y = lat, group = group)) +
geom_polygon(fill = "cornsilk4", col = "cornsilk") +
geom_polygon(data=counties,color="black",fill="lightgoldenrod") +
geom_polygon(data = ranas4, fill = "green", alpha = 0.5) +
theme_void() +
theme(panel.background = element_rect(fill = "cornsilk")) +
coord_map("albers",lat=90,lat1=80)
# Tried to trim data outside California (Mexico data) with no success.
#I tried:
ranas2 <- subset(ranas,ORIGIN !=1)
but it doesn't have any effect or subsets anything. 回答 1
Stack Overflow用户
发布于 2022-05-05 04:36:00
细分您的空间数据将没有多大用处,因为它的两个特性(行)都不完全在加利福尼亚:
## plot features by origin:
library(ggplot2)
library(sf)
my_shp <- read_sf(dsn = "path_to_shp_directory")
my_shp %>%
ggplot() +
geom_sf() +
facet_wrap(~ ORIGIN)您仍然可以使用带有加利福尼亚边界的(剪辑)功能.:
## make sure both geometries have the same CRS,
## if necessary, by st_transform(your_epsg_code)
my_shp_cropped <- st_crop(my_shp, cali_map)..。但是请注意,这不会重新计算底层的青蛙数据(例如,只在加利福尼亚计算青蛙数量)。
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/72121806
复制相关文章
相似问题

