对字符串数据的部分进行热编码。
ACN58946.1|FEATURES|NCBI|macrolide-lincosamide-streptogramin|macB|antibiotic靶protection|0\sMAEPVLSVKDLDIRFTTPDGNVHAVKKVSFDIAPGECLGVVGESGSGKSQLFMACIGLLAGNGKATGSVTYRGQELLGQPAAKLNAIRGAKITMIFQDPLTSLTPHMRIGDQIVESLRTHSKLSKGEAEKRAIQALELVRIPEAKRRMRQYPHELSGGMRQRVMIAMATACGPDLLIADEPTTALDVTVQAQILDIMRDLRKELGTSIALISHDMGVIASICDRVQVMRYGEFVETGPADDIFYHPQHPYTRMLLEAMPRIDQPVREGRAALKPLAPQEARTTLLEVDNVKVHFPIQMGGVFFGKYKPLRAVDGVSFTLHQGETIGIVGESGCGKSTLARAVLELLPKTTGGVVWMGRDLGALPPAELRRARKDFQIVFQDPLASLDPRMTIGQSIAEPLQSLEPELSKHEVQSRVRAIMEKVGLDPDWINRYPHEFSGGQNQRVGIARAMILKPKLIVCDEAVSALDVSIQAQIVDLILSLQAEFGMSIIFISHDLSVVRQVSHRVMVLYLGRVVELASRDAIYEDARHPYTKALISAVTVPDPRAERLKKRRELPGELPSPLDTRSALMFLKSKRIDDPDAEQYVPKLIEVAPGHFVAEHDPFEVVEMTG\e>ACN58871.1|FEATURES|NCBI|macrolide-lincosamide-streptogramin|macB|antibiotic efflux|0\sMADYLLEMKNIVKEFGGVRALNGIDIKLKAGECAGLCGENGAGKSTLMKVLSAVYPHGTWDGEILWDGKPLRAQSIRETEAAGIVIIHQELMLVPELSVAENIFLGNEIKLPGGRMDYAAMNRRAEELLAELDIRDVNVVLPVKQYGGGYQQLIEIAKALNKNARLLILDEPSSSLTASEIKVLLRIIHSLKAKGVTCVYISHKLDEVADICDTIVVIRDGQHIATTPMADMNIERIIAQMVGREMNQLYPERSHVPGEVIFEARNVSCYDADNPQRKRVDNISFKLRKGEILGIAGLVGAGRTELVSALFGAYPGPSEAEVWLNGVKLDTRTPLKAIRAGLAMVPEDRKQHGIVPDLGVGHNMTLAVLNDFVRATRIDQQAELATIHKEIKSVKLKTATPFLPITSLSGGNQQKAVLSKMLLTKPKILILDEPTRGVDVGAKFEIYQLMFDLAAQGMSIIMVSSELAEVLGISDRVLVVGEGKLRGDFVNDNLSQETVLAAALDHTQPALH\e>ACN58991.1|FEATURES|NCBI|multidrug|cmeB|antibiotic efflux|0\sMKNDRGEMVPFSAFMTIKKKQGANEINRYNMYNTAAIRGGPATGYSSGEAIKAVQEVAAKNLPNGFDIDWAALSYDETRRGNEAVYIFLIVLAFVYLVLAAQYESFIIPLAVVFSLPAGVFGSFLLIKGMGLANDIYAQVGLVMLVGLLGKNAVLIVEFAVQKQQQGATVFEAAIEGARVRFRPILMTSFAFIAGLIPLVFAHGAGAIGNKTIGSSALGGMFFGTVFGVIVVPGLYYVFGSWAEGRKLIRGEDHDPLTENLVHQMDNFPQSDDK\e>ACN58776.1|FEATURES|NCBI|macrolide-lincosamide-streptogramin|macA|antibiotic efflux|0\sMGNLPRPTLSPSLSGIRPTMNRETTTRVDSSTPAARLGMRVPSTSRAALVGVAALVVILGGWYGIKRWRAHVASEGQYIFAAIQKGDIEDLVTATGSLQPRDYVDVGAQVSGQLDKILVEVGSDVKEGDLLAEIDADVAAARVDASRAQLRSQQAQLVQQQANLTKAERDLTRQQNLMKEDATTAEQVQNAETTLDTTKAQINALKAQMEQLRASMRVDESNLNYTKILAPMSGTVVSISAKQGQTLNTNQQAPTILRIADLSTMTVQTQVSEADVSKLRSGMQAYFTTLGSAGKRWYGQLKKIEPTPTVTNNVVLYNALFEVPNDNKQLLPQMTAQVFFVAAAAHDVLVVPMSAVSLQRTPPGGIPNAAAAQAAGARGAGAQGAGAQGAQGASAQGAGAQSGQGGQGAAALTPEQIARREARRQQRMQSNGGSATGGAIEGGPPRGGFGASMAARGPRHATVRVQAADGKIEERQITIGVTNRVHAEVLSGLKEGERVVAGTKEPEKAPATAGGQQGAGGQRNNIGGFPGGGLGGGFGR\e
我现在正在研究蛋白质序列。我有这些字符串数据,我想要转换成一个热编码部分。我想要转换的部分在'\s‘之后开始,在'\e’之前结束,然后对整个字符串数据进行转换。由于I有数千个数据集,使用纯Python代码似乎是不可能的,因为完成这个过程需要很长时间。针对这个问题有机器学习库吗?
谢谢你提前提供帮助!
回答 2
Stack Overflow用户
发布于 2022-05-04 16:18:20
我建议使用斯克勒恩的单热编码器。它应该以最大的效率对数据进行编码。
如果您想在Python中优化其他ML任务,我建议查看丹瑟尔流/喀拉斯、PyTorch 科学知识-学习(SKLearn)和ScyPi。当然,Numpy也是必不可少的。这些工具功能强大,优化程度很高,并且有大量的资源可以帮助您解决问题。它们应该可以帮助您解决几乎所有遇到的ML问题:)
Stack Overflow用户
发布于 2022-05-05 01:36:30
为此您可以使用熊猫。
import pandas as pd
import io
data = "ACN58946.1|FEATURES|NCBI|macrolide-lincosamide-streptogramin|macB|antibiotic target protection|0\sMAEPVLSVKDLDIRFTTPDGNVHAVKKVSFDIAPGECLGVVGESGSGKSQLFMACIGLLAGNGKATGSVTYRGQELLGQPAAKLNAIRGAKITMIFQDPLTSLTPHMRIGDQIVESLRTHSKLSKGEAEKRAIQALELVRIPEAKRRMRQYPHELSGGMRQRVMIAMATACGPDLLIADEPTTALDVTVQAQILDIMRDLRKELGTSIALISHDMGVIASICDRVQVMRYGEFVETGPADDIFYHPQHPYTRMLLEAMPRIDQPVREGRAALKPLAPQEARTTLLEVDNVKVHFPIQMGGVFFGKYKPLRAVDGVSFTLHQGETIGIVGESGCGKSTLARAVLELLPKTTGGVVWMGRDLGALPPAELRRARKDFQIVFQDPLASLDPRMTIGQSIAEPLQSLEPELSKHEVQSRVRAIMEKVGLDPDWINRYPHEFSGGQNQRVGIARAMILKPKLIVCDEAVSALDVSIQAQIVDLILSLQAEFGMSIIFISHDLSVVRQVSHRVMVLYLGRVVELASRDAIYEDARHPYTKALISAVTVPDPRAERLKKRRELPGELPSPLDTRSALMFLKSKRIDDPDAEQYVPKLIEVAPGHFVAEHDPFEVVEMTG\e>ACN58871.1|FEATURES|NCBI|macrolide-lincosamide-streptogramin|macB|antibiotic efflux|0\sMADYLLEMKNIVKEFGGVRALNGIDIKLKAGECAGLCGENGAGKSTLMKVLSAVYPHGTWDGEILWDGKPLRAQSIRETEAAGIVIIHQELMLVPELSVAENIFLGNEIKLPGGRMDYAAMNRRAEELLAELDIRDVNVVLPVKQYGGGYQQLIEIAKALNKNARLLILDEPSSSLTASEIKVLLRIIHSLKAKGVTCVYISHKLDEVADICDTIVVIRDGQHIATTPMADMNIERIIAQMVGREMNQLYPERSHVPGEVIFEARNVSCYDADNPQRKRVDNISFKLRKGEILGIAGLVGAGRTELVSALFGAYPGPSEAEVWLNGVKLDTRTPLKAIRAGLAMVPEDRKQHGIVPDLGVGHNMTLAVLNDFVRATRIDQQAELATIHKEIKSVKLKTATPFLPITSLSGGNQQKAVLSKMLLTKPKILILDEPTRGVDVGAKFEIYQLMFDLAAQGMSIIMVSSELAEVLGISDRVLVVGEGKLRGDFVNDNLSQETVLAAALDHTQPALH\e>ACN58991.1|FEATURES|NCBI|multidrug|cmeB|antibiotic efflux|0\sMKNDRGEMVPFSAFMTIKKKQGANEINRYNMYNTAAIRGGPATGYSSGEAIKAVQEVAAKNLPNGFDIDWAALSYDETRRGNEAVYIFLIVLAFVYLVLAAQYESFIIPLAVVFSLPAGVFGSFLLIKGMGLANDIYAQVGLVMLVGLLGKNAVLIVEFAVQKQQQGATVFEAAIEGARVRFRPILMTSFAFIAGLIPLVFAHGAGAIGNKTIGSSALGGMFFGTVFGVIVVPGLYYVFGSWAEGRKLIRGEDHDPLTENLVHQMDNFPQSDDK\e>ACN58776.1|FEATURES|NCBI|macrolide-lincosamide-streptogramin|macA|antibiotic efflux|0\sMGNLPRPTLSPSLSGIRPTMNRETTTRVDSSTPAARLGMRVPSTSRAALVGVAALVVILGGWYGIKRWRAHVASEGQYIFAAIQKGDIEDLVTATGSLQPRDYVDVGAQVSGQLDKILVEVGSDVKEGDLLAEIDADVAAARVDASRAQLRSQQAQLVQQQANLTKAERDLTRQQNLMKEDATTAEQVQNAETTLDTTKAQINALKAQMEQLRASMRVDESNLNYTKILAPMSGTVVSISAKQGQTLNTNQQAPTILRIADLSTMTVQTQVSEADVSKLRSGMQAYFTTLGSAGKRWYGQLKKIEPTPTVTNNVVLYNALFEVPNDNKQLLPQMTAQVFFVAAAAHDVLVVPMSAVSLQRTPPGGIPNAAAAQAAGARGAGAQGAGAQGAQGASAQGAGAQSGQGGQGAAALTPEQIARREARRQQRMQSNGGSATGGAIEGGPPRGGFGASMAARGPRHATVRVQAADGKIEERQITIGVTNRVHAEVLSGLKEGERVVAGTKEPEKAPATAGGQQGAGGQRNNIGGFPGGGLGGGFGR\e"
data = data.replace('\e>', '\r')
with io.BytesIO(data.encode('utf8')) as binary_file:
with io.TextIOWrapper(binary_file, encoding='utf8') as file_obj:
df = pd.read_table(file_obj, sep="|", header=None, )
df[6] = df[6].str.replace('0\s', '', regex=False)像这样给你买一张数据文件:
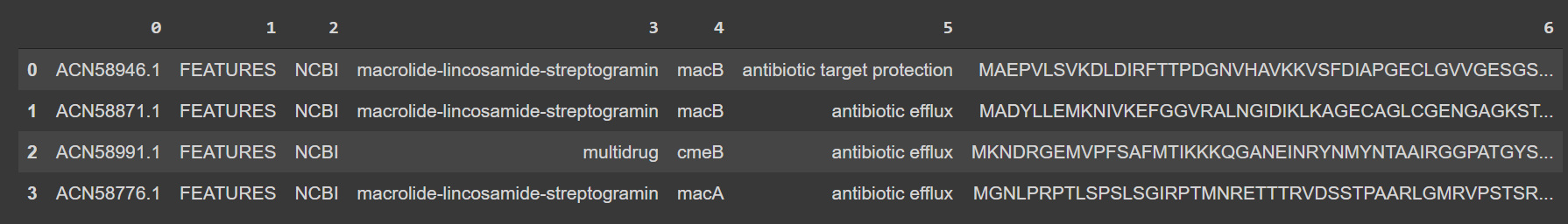
之后,您可以使用sklearn.preprocessing.OneHotEncoder或假人编码您的蛋白质序列。我在这里展示了熊猫的版本:
df = pd.get_dummies(data=df, columns=[6])这是一个例子,如果您的数据在一个变量中可用。如果是文本文件,则不需要io方法,应该用加载txt文件来替换。还要注意,应该用换行符(例如"\r“或"\n”)替换"\e>“来分离样本。
https://stackoverflow.com/questions/72114227
复制相似问题

