使用rstatix::wilcox_test()、rstatix::add_xy_position()和ggpubr::stat_pvalue_manual(),p值位置被错误地定义为零
使用rstatix::wilcox_test()、rstatix::add_xy_position()和ggpubr::stat_pvalue_manual(),p值位置被错误地定义为零
提问于 2022-04-15 20:59:01
在对我的数据执行rstatix::wilcox_test()之后,我自动添加了y.position值,以便使用rstatix::add_xy_position()绘制p值。然后,我绘制了数据并使用ggpubr::stat_pvalue_manual()添加了p值。问题是,y.position值被错误地定义为零,导致所绘制的p值和括号geom出现在y-轴底部的零位置,而此时它应该出现在y-轴顶部附近。rstatix的文档表明,按max默认设置计算的y.position值应该等于max(data$Response) = 0.00008556344,而不是零。我是不是不正确地使用这些函数?如果不是,如何才能使p值和括号geom出现在对应于最大值的y轴数据值上?
library(tidyverse)
library(rstatix)
library(ggpubr)
# minimal data example
data <- tibble::tribble(
~Group, ~Response,
"Treatment", 3.210486e-06,
"Control", 4.006825e-06,
"Treatment", 4.350836e-06,
"Control", 4.216934e-06,
"Treatment", 4.415194e-06,
"Control", 1.0260606e-05,
"Treatment", 1.111064e-06,
"Control", 1.0779088e-05,
"Treatment", 3.57185e-07,
"Control", 1.139097e-06,
"Treatment", 0,
"Control", 2.31074e-07,
"Treatment", 5.78956e-07,
"Control", 4.371157e-06,
"Treatment", 6.5825e-08,
"Control", 9.587202e-06,
"Treatment", 2.65383e-07,
"Control", 3.57337e-06,
"Treatment", 7.14146e-07,
"Control", 3.868605e-06,
"Treatment", 1.2213951e-05,
"Control", 6.936899e-06,
"Treatment", 4.71707e-07,
"Control", 5.5957173e-05,
"Treatment", 1.265942e-06,
"Control", 8.5563441e-05,
"Treatment", 0,
"Control", 0,
"Treatment", 0,
"Control", 2.1306289e-05,
"Treatment", 5.2055e-07,
"Control", 1.8420094e-05
)
# performing Wilcoxon test and adding 'y.position' for the p value
wilcoxon_result <- data %>%
wilcox_test(Response ~ Group) %>%
add_xy_position("Group")
# the 'y.position' is incorrectly defined as zero.
wilcoxon_result
#> # A tibble: 1 × 11
#> .y. group1 group2 n1 n2 statistic p y.position groups xmin
#> <chr> <chr> <chr> <int> <int> <dbl> <dbl> <dbl> <name> <dbl>
#> 1 Response Control Treatm… 16 16 206. 0.00326 0 <chr> 1
#> # … with 1 more variable: xmax <dbl>
# the p value and bracket geom are plotted at zero.
ggplot(data, aes(Group, Response)) +
stat_summary(geom = "errorbar", fun.data = mean_cl_boot, width = 0.25) +
stat_summary(geom = "crossbar", fun = mean, fun.min = mean, fun.max = mean, fatten = FALSE, width = 0.5) +
stat_pvalue_manual(wilcoxon_result, "P = {p}", size = 18 / ggplot2::.pt, tip.length = 0, bracket.size = 2 / ggplot2::.stroke) +
theme_pubr(base_size = 18, x.text.angle = 45)

回答 1
Stack Overflow用户
回答已采纳
发布于 2022-04-15 21:25:18
似乎你的y位被四舍五入为零,因为你处理的是非常小的数字。如果将响应变量乘以100万,则得到相同的p值,然后可以将y位置除以使其位于任何您想要的位置:
# performing Wilcoxon test and adding 'y.position' for the p value
wilcoxon_result <- data %>%
mutate(Response = Response * 1e6) %>%
wilcox_test(Response ~ Group) %>%
add_xy_position("Group") %>%
mutate(y.position = y.position /3e6)这意味着:
wilcoxon_result
#> # A tibble: 1 x 11
#> .y. group1 group2 n1 n2 statistic p y.position groups xmin xmax
#> <chr> <chr> <chr> <int> <int> <dbl> <dbl> <dbl> <named list> <dbl> <dbl>
#> Response Control Treatment 16 16 206. 0.00326 0.0000315 <chr [2]> 1 2情节是这样的:
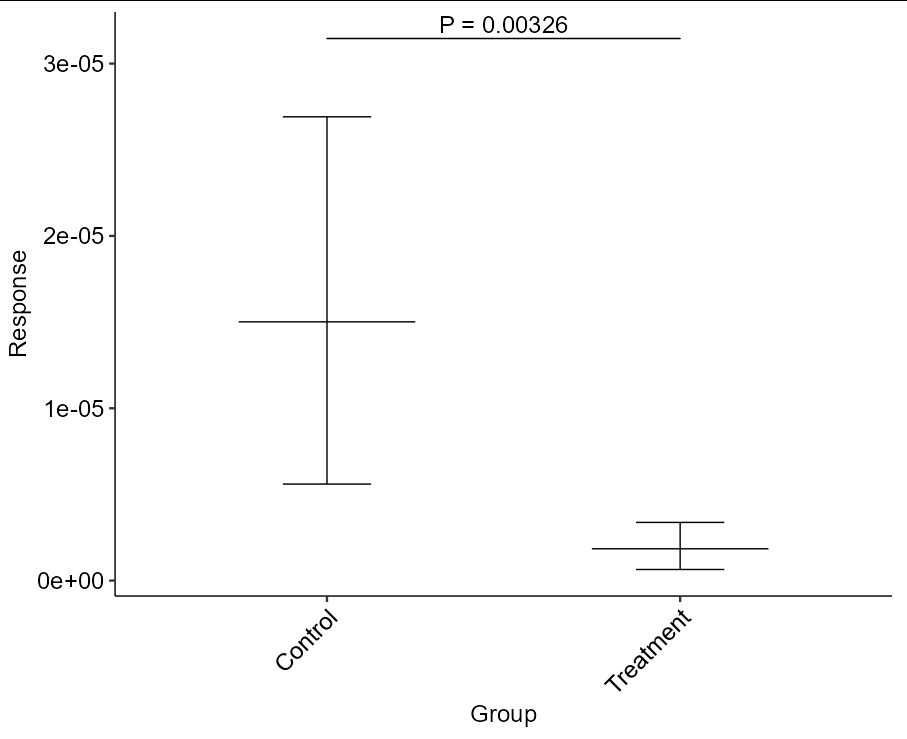
ggplot(data, aes(Group, Response)) +
stat_summary(geom = "errorbar", fun.data = mean_cl_boot, width = 0.25) +
stat_summary(geom = "crossbar", fun = mean, fun.min = mean,
fun.max = mean, fatten = FALSE, width = 0.5) +
stat_pvalue_manual(wilcoxon_result, "P = {p}",
size = 18 / ggplot2::.pt, tip.length = 0,
bracket.size = 2 / ggplot2::.stroke) +
theme_pubr(base_size = 18, x.text.angle = 45)
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/71888899
复制相关文章
相似问题

