日志变量转换后GGplot2在check_aesthetics中的错误
日志变量转换后GGplot2在check_aesthetics中的错误
提问于 2022-04-12 14:10:50
我试图用相同均值和标准差的理论正态分布来绘制密度直方图。
由于数据是倾斜的,所以我尝试了日志(1+x)转换:
knitr::opts_chunk$set(echo = TRUE)
library("tidyverse", "here", "magrittr")
dlf<-here::here("data/DownloadFestival(No Outlier).dat") %>% readr::read_delim(col_names = TRUE)day2log<-na.omit(log1p(dlf$day2))
day2log<-as.numeric(day2log)
glimpse(day2log)
hist.day2log<-ggplot2::ggplot(dlf, aes(day2log)) +
geom_histogram(aes(y=..density..), colour = "#76b81b", fill = "blue" ) +
stat_function(fun = dnorm, args = list(mean = mean(day2log), sd = sd(day2log)), colour = "black", size = 2) +
labs( x = "Hygiene score day 2log", y = "Density day2log") +
theme_minimal()
hist.day2log我得到的错误如下:
num [1:264] 0.854 0.88 0.077 0.365 0.182 ...
Error in `check_aesthetics()`:
! Aesthetics must be either length 1 or the same as the data (810): x
Backtrace:
1. base `<fn>`(x)
2. ggplot2:::print.ggplot(x)
4. ggplot2:::ggplot_build.ggplot(x)
5. ggplot2 by_layer(function(l, d) l$compute_aesthetics(d, plot))
6. ggplot2 f(l = layers[[i]], d = data[[i]])
7. l$compute_aesthetics(d, plot)
8. ggplot2 f(..., self = self)
9. ggplot2:::check_aesthetics(evaled, n)
Error in check_aesthetics(evaled, n) :我试着检查day2log变量,它看起来很好:
glimpse(day2log)
num [1:264] 0.854 0.88 0.077 0.365 0.182 ...回答 1
Stack Overflow用户
回答已采纳
发布于 2022-04-12 14:22:00
您已经从NA中删除了day2值,因此向量day2log现在比dlf短,dlf是要传递到ggplot的数据帧。由于ggplot期望任何外部向量(即不是传递的数据帧中的列的向量)与数据帧的长度相同,或者长度为1,则会出现错误。
要解决这个问题,直接在day2上进行计算。您可以显式删除需要删除的NA值,例如在mean和sd中。
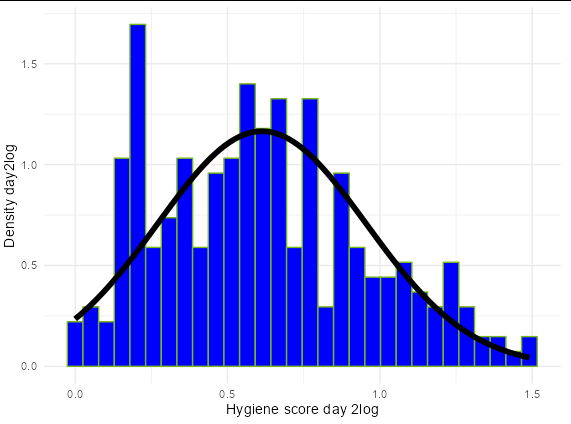
ggplot(dlf, aes(log1p(day2))) +
geom_histogram(aes(y=..density..), colour = "#76b81b", fill = "blue" ) +
stat_function(fun = dnorm,
args = list(mean = mean(log1p(dlf$day2), na.rm = TRUE),
sd = sd(log1p(dlf$day2), na.rm = TRUE)),
colour = "black", size = 2) +
labs( x = "Hygiene score day 2log", y = "Density day2log") +
theme_minimal()
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/71844221
复制相关文章
相似问题

