使用diff()函数创建条形图,以确定下一个事件的生存变化
使用diff()函数创建条形图,以确定下一个事件的生存变化
提问于 2022-03-27 16:29:34
我有一个生存数据集df_survival,它有四列Date、Tank、Feed_Group和Census。
下面是创建数据集的代码:
Date <- c("2020-10-15", "2020-10-15", "2020-10-15", "2020-10-15", "2020-10-15", "2020-10-15","2020-10-15", "2020-10-15","2020-12-05","2020-12-05","2020-12-05","2020-12-05","2020-12-05","2020-12-05","2020-12-05","2020-12-05","2021-01-29","2021-01-29","2021-01-29","2021-01-29","2021-01-29","2021-01-29","2021-01-29","2021-03-13","2021-03-13","2021-03-13","2021-03-13","2021-03-13","2021-03-13","2021-03-13","2021-03-13","2021-04-30","2021-04-30","2021-04-30","2021-04-30","2021-04-30","2021-04-30","2021-04-30","2021-04-30","2021-04-30", "2021-07-06", "2021-07-06", "2021-07-06", "2021-07-06", "2021-07-06", "2021-07-06", "2021-07-06", "2021-07-06")
Tank <- c(1,2,3,4,5,6,7,8,1,2,3,4,5,6,7,8,1,2,3,4,5,6,7,8,1,2,3,4,5,6,7,8,1,2,3,4,5,6,7,8,1,2,3,4,5,6,7,8)
Feed_Group <- c("Gemma","Gemma","Gemma","Gemma","Mysis","Mysis","Mysis","Mysis","Gemma","Gemma","Gemma","Gemma","Mysis","Mysis","Mysis","Mysis","Gemma","Gemma","Gemma","Gemma","Mysis","Mysis","Mysis","Mysis","Gemma","Gemma","Gemma","Gemma","Mysis","Mysis","Mysis","Mysis","Gemma","Gemma","Gemma","Gemma","Mysis","Mysis","Mysis","Mysis","Gemma","Gemma","Gemma","Gemma","Mysis","Mysis","Mysis","Mysis")
Census <- c(20,20,20,20,20,20,20,20,15,17,14,10,14,20,16,16,13,16,12,10,14,20,16,16,13,15,11,10,14,20,16,16,13,15,10,9,14,20,15,16,13,14,10,8,14,19,15,15)
df_survival <- data.frame(Date,Tank,Feed_Group,Census)我创建了这个可视化系统,它显示了随着时间的推移,每个群体的平均人口普查结果,并带有标准错误。
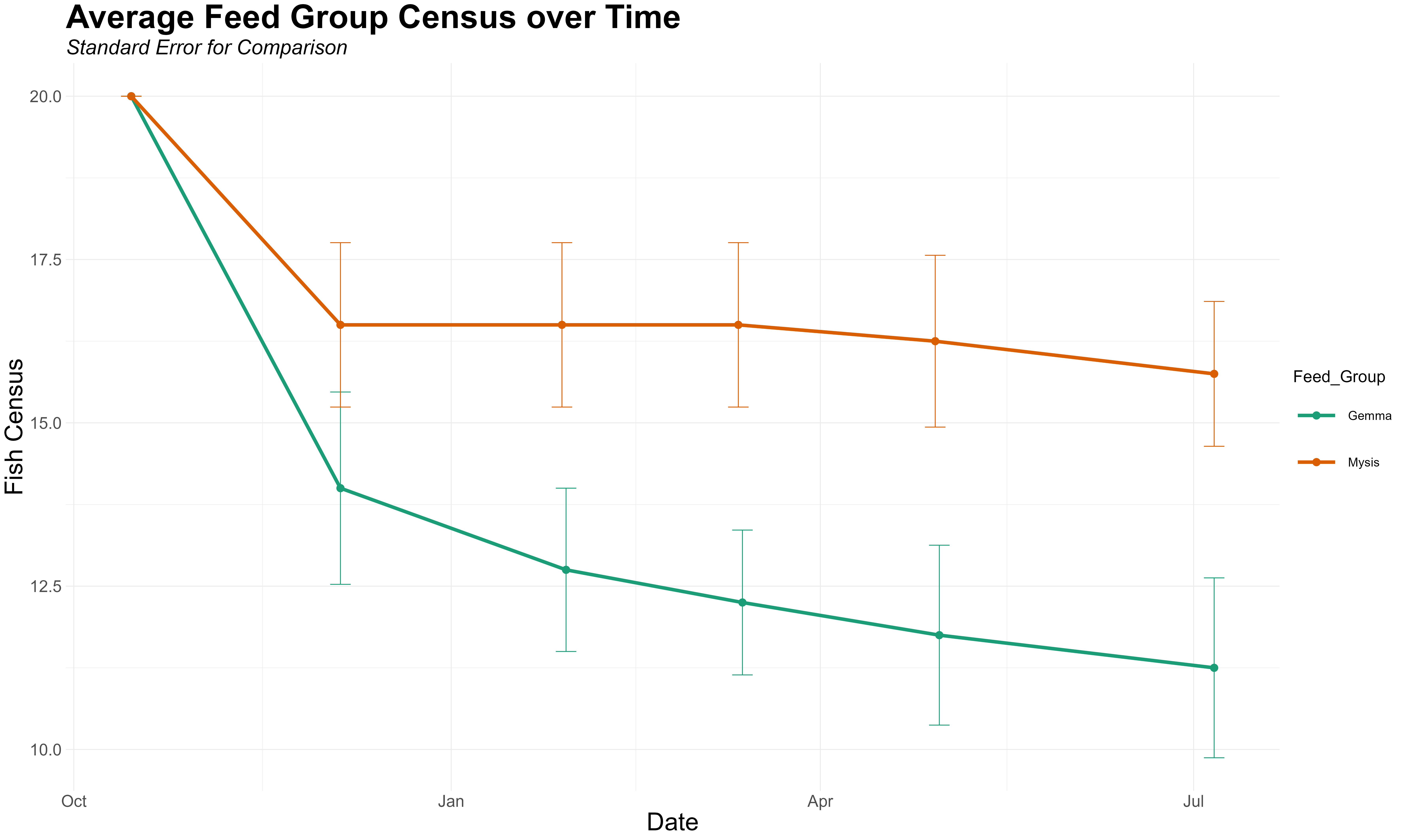
现在,我想要创建一个条形图,显示在一个事件到下一个事件中有多少动物死亡。因此,从事件1到事件2,引理组有6只动物减少,从事件2减少到事件3,1.25只动物减少。
我可以像下面这样使用diff()函数来创建一个tibble,它为我提供了一个与我想要的完全一样的Feed_Group和Census_Dead列。
df_survival %>%
group_by(Feed_Group,Date) %>%
summarise(Census = mean(Census)) %>%
summarise(Census_Dead = diff(Census))但是,我很可能会丢失Date列,因为第一个点没有0的差异,这是有道理的。不幸的是,这使得作图变得很困难。
我知道我可以创建data.frame并手动将这些数字与相应的日期放在一起,因为数据集太小了,但是我很好奇是否有可能操作当前的数据以获得如下内容:
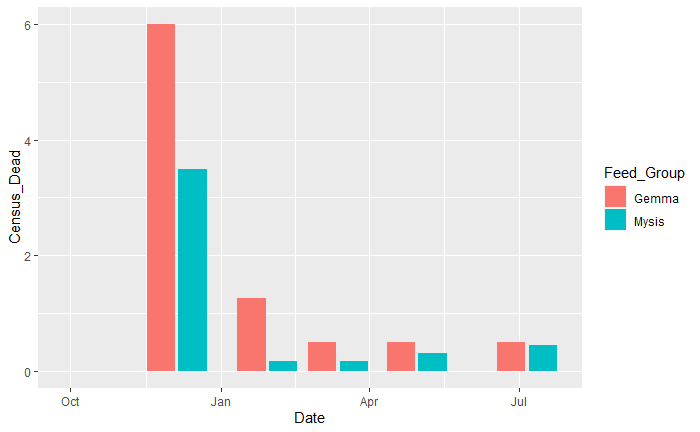
回答 1
Stack Overflow用户
回答已采纳
发布于 2022-03-28 09:22:37
可以使用dplyr::summarize为每个组获取更少的行(而不仅仅是聚合到一行)。或者,您可以使用dplyr::group_modify。
df_survival |>
group_by(Date, Feed_Group) |>
summarize(Census = mean(Census)) |>
ungroup() |>
arrange(Date) |>
group_by(Feed_Group) |>
summarize(Census_Dead = -diff(Census),
Date = Date[-1]) |>
ggplot(aes(y = Census_Dead, x= Date)) +
geom_bar(aes(fill = Feed_Group), position = "dodge", stat = "identity")
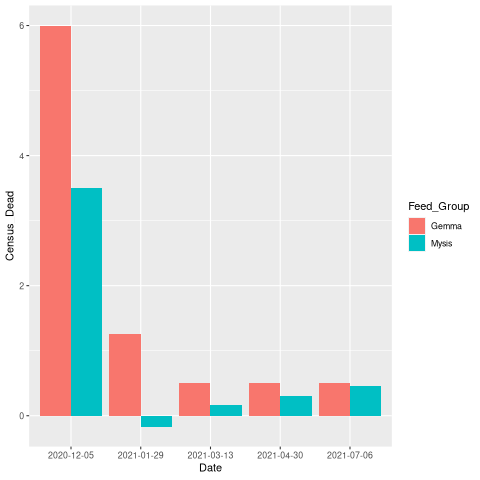
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/71638165
复制相关文章
相似问题

