生成所有给定大小的有向图,直至同构
我正在尝试生成所有具有给定节点数的有向图,直到图同构,这样我就可以将它们提供给另一个Python程序。下面是一个使用NetworkX的简单参考实现,我想加快它的速度:
from itertools import combinations, product
import networkx as nx
def generate_digraphs(n):
graphs_so_far = list()
nodes = list(range(n))
possible_edges = [(i, j) for i, j in product(nodes, nodes) if i != j]
for edge_mask in product([True, False], repeat=len(possible_edges)):
edges = [edge for include, edge in zip(edge_mask, possible_edges) if include]
g = nx.DiGraph()
g.add_nodes_from(nodes)
g.add_edges_from(edges)
if not any(nx.is_isomorphic(g_before, g) for g_before in graphs_so_far):
graphs_so_far.append(g)
return graphs_so_far
assert len(generate_digraphs(1)) == 1
assert len(generate_digraphs(2)) == 3
assert len(generate_digraphs(3)) == 16这类图的数量似乎增长很快,由这个OEIS序列给出。我正在寻找一种解决方案,能够在合理的时间内生成最多7个节点的所有图(总计约10亿个图)。
将图表示为NetworkX对象并不是很重要;例如,表示具有邻接列表的图或使用不同的库对我有好处。
回答 3
Stack Overflow用户
发布于 2022-04-01 03:37:38
有一个有用的想法,我从布兰登·麦凯( Brendan )的论文“无异形无穷尽世代”(虽然我相信它早于那篇论文)中学到的。
其思想是,我们可以将同构类组织成一棵树,其中带空图的单例类是根,每个具有n>0节点的图的类都有一个具有n个−1节点的图的父类。为了枚举n>0节点的图的同构类,枚举具有n个−1节点的图的同构类,并对每个此类类以所有可能的方式将其表示扩展到n个节点,并滤除那些实际上不是子节点的同构类。
下面的Python代码使用一个基本但非平凡的图同构子例程来实现这一思想。N=6只需几分钟,而n= 7则需要几天的时间估计。对于额外的速度,请将其移植到C++,并可能找到更好的算法来处理置换组(可能在TAoCP中,尽管大多数图没有对称性,因此不清楚收益有多大)。
import cProfile
import collections
import itertools
import random
# Returns labels approximating the orbits of graph. Two nodes in the same orbit
# have the same label, but two nodes in different orbits don't necessarily have
# different labels.
def invariant_labels(graph, n):
labels = [1] * n
for r in range(2):
incoming = [0] * n
outgoing = [0] * n
for i, j in graph:
incoming[j] += labels[i]
outgoing[i] += labels[j]
for i in range(n):
labels[i] = hash((incoming[i], outgoing[i]))
return labels
# Returns the inverse of perm.
def inverse_permutation(perm):
n = len(perm)
inverse = [None] * n
for i in range(n):
inverse[perm[i]] = i
return inverse
# Returns the permutation that sorts by label.
def label_sorting_permutation(labels):
n = len(labels)
return inverse_permutation(sorted(range(n), key=lambda i: labels[i]))
# Returns the graph where node i becomes perm[i] .
def permuted_graph(perm, graph):
perm_graph = [(perm[i], perm[j]) for (i, j) in graph]
perm_graph.sort()
return perm_graph
# Yields each permutation generated by swaps of two consecutive nodes with the
# same label.
def label_stabilizer(labels):
n = len(labels)
factors = (
itertools.permutations(block)
for (_, block) in itertools.groupby(range(n), key=lambda i: labels[i])
)
for subperms in itertools.product(*factors):
yield [i for subperm in subperms for i in subperm]
# Returns the canonical labeled graph isomorphic to graph.
def canonical_graph(graph, n):
labels = invariant_labels(graph, n)
sorting_perm = label_sorting_permutation(labels)
graph = permuted_graph(sorting_perm, graph)
labels.sort()
return max(
(permuted_graph(perm, graph), perm[sorting_perm[n - 1]])
for perm in label_stabilizer(labels)
)
# Returns the list of permutations that stabilize graph.
def graph_stabilizer(graph, n):
return [
perm
for perm in label_stabilizer(invariant_labels(graph, n))
if permuted_graph(perm, graph) == graph
]
# Yields the subsets of range(n) .
def power_set(n):
for r in range(n + 1):
for s in itertools.combinations(range(n), r):
yield list(s)
# Returns the set where i becomes perm[i] .
def permuted_set(perm, s):
perm_s = [perm[i] for i in s]
perm_s.sort()
return perm_s
# If s is canonical, returns the list of permutations in group that stabilize s.
# Otherwise, returns None.
def set_stabilizer(s, group):
stabilizer = []
for perm in group:
perm_s = permuted_set(perm, s)
if perm_s < s:
return None
if perm_s == s:
stabilizer.append(perm)
return stabilizer
# Yields one representative of each isomorphism class.
def enumerate_graphs(n):
assert 0 <= n
if 0 == n:
yield []
return
for subgraph in enumerate_graphs(n - 1):
sub_stab = graph_stabilizer(subgraph, n - 1)
for incoming in power_set(n - 1):
in_stab = set_stabilizer(incoming, sub_stab)
if not in_stab:
continue
for outgoing in power_set(n - 1):
out_stab = set_stabilizer(outgoing, in_stab)
if not out_stab:
continue
graph, i_star = canonical_graph(
subgraph
+ [(i, n - 1) for i in incoming]
+ [(n - 1, j) for j in outgoing],
n,
)
if i_star == n - 1:
yield graph
def test():
print(sum(1 for graph in enumerate_graphs(5)))
cProfile.run("test()")Stack Overflow用户
发布于 2022-03-28 14:15:17
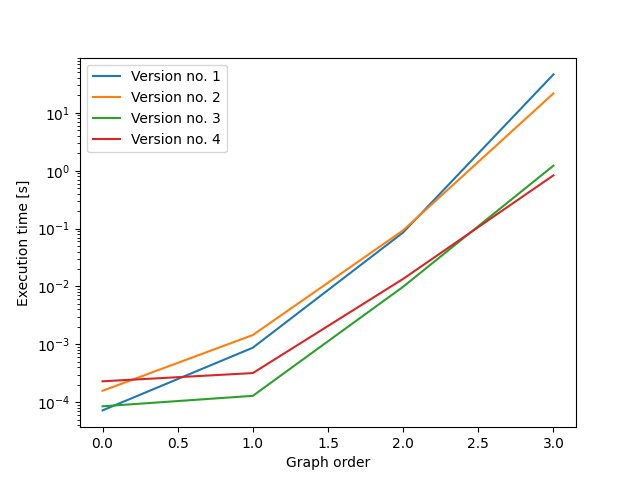
98-99%的计算时间用于同构测试,因此游戏的名称是为了减少必要的测试次数。在这里,我以批方式创建了图,因此只有在批内才需要测试图的同构。
在第一个变体(以下版本2)中,批处理中的所有图都有相同的边数。这导致了运行时间的可理解但适度的改进(对于大小为4的图,速度是4的2.5倍,对于较大的图,则有更大的速度增长)。
在第二个变体(以下版本3)中,批处理中的所有图都有相同的出度序列。这将大大提高运行时间(对于大小为4的图形,运行速度是4的35倍,而对于较大的图,则提高了更大的速度)。
在第三个变体(以下版本4)中,批处理中的所有图都有相同的出度序列。此外,在一批图中,所有图都是按内次序列排序的.这使速度比第3版略有提高(尺寸为4的图形比第4版的速度快1.3倍;尺寸5的图形的速度快2.1倍)。
#!/usr/bin/env python
"""
Efficient motif generation.
"""
import numpy as np
import matplotlib.pyplot as plt
import networkx as nx
from timeit import timeit
from itertools import combinations, product, chain, combinations_with_replacement
# for profiling with kernprof/line_profiler
try:
profile
except NameError:
profile = lambda x: x
@profile
def version_1(n):
"""Original implementation by @hilberts_drinking_problem"""
graphs_so_far = list()
nodes = list(range(n))
possible_edges = [(i, j) for i, j in product(nodes, nodes) if i != j]
for edge_mask in product([True, False], repeat=len(possible_edges)):
edges = [edge for include, edge in zip(edge_mask, possible_edges) if include]
g = nx.DiGraph()
g.add_nodes_from(nodes)
g.add_edges_from(edges)
if not any(nx.is_isomorphic(g_before, g) for g_before in graphs_so_far):
graphs_so_far.append(g)
return graphs_so_far
@profile
def version_2(n):
"""Creates graphs in batches, where each batch contains graphs with
the same number of edges. Only graphs within a batch have to be tested
for isomorphisms."""
graphs_so_far = list()
nodes = list(range(n))
possible_edges = [(i, j) for i, j in product(nodes, nodes) if i != j]
for ii in range(len(possible_edges)+1):
tmp = []
for edges in combinations(possible_edges, ii):
g = nx.from_edgelist(edges, create_using=nx.DiGraph)
if not any(nx.is_isomorphic(g_before, g) for g_before in tmp):
tmp.append(g)
graphs_so_far.extend(tmp)
return graphs_so_far
@profile
def version_3(n):
"""Creates graphs in batches, where each batch contains graphs with
the same out-degree sequence. Only graphs within a batch have to be tested
for isomorphisms."""
graphs_so_far = list()
outdegree_sequences_so_far = list()
for outdegree_sequence in product(*[range(n) for _ in range(n)]):
# skip degree sequences which we have already seen as the resulting graphs will be isomorphic
if sorted(outdegree_sequence) not in outdegree_sequences_so_far:
tmp = []
for edges in generate_graphs(outdegree_sequence):
g = nx.from_edgelist(edges, create_using=nx.DiGraph)
if not any(nx.is_isomorphic(g_before, g) for g_before in tmp):
tmp.append(g)
graphs_so_far.extend(tmp)
outdegree_sequences_so_far.append(sorted(outdegree_sequence))
return graphs_so_far
def generate_graphs(outdegree_sequence):
"""Generates all directed graphs with a given out-degree sequence."""
for edges in product(*[generate_edges(node, degree, len(outdegree_sequence)) \
for node, degree in enumerate(outdegree_sequence)]):
yield(list(chain(*edges)))
def generate_edges(node, outdegree, total_nodes):
"""Generates all edges for a given node with a given out-degree and a given graph size."""
for targets in combinations(set(range(total_nodes)) - {node}, outdegree):
yield([(node, target) for target in targets])
@profile
def version_4(n):
"""Creates graphs in batches, where each batch contains graphs with
the same out-degree sequence. Within a batch, graphs are sorted
by in-degree sequence, such that only graphs with the same
in-degree sequence have to be tested for isomorphism.
"""
graphs_so_far = list()
for outdegree_sequence in combinations_with_replacement(range(n), n):
tmp = dict()
for edges in generate_graphs(outdegree_sequence):
g = nx.from_edgelist(edges, create_using=nx.DiGraph)
indegree_sequence = tuple(sorted(degree for _, degree in g.in_degree()))
if indegree_sequence in tmp:
if not any(nx.is_isomorphic(g_before, g) for g_before in tmp[indegree_sequence]):
tmp[indegree_sequence].append(g)
else:
tmp[indegree_sequence] = [g]
for graphs in tmp.values():
graphs_so_far.extend(graphs)
return graphs_so_far
if __name__ == '__main__':
order = range(1, 5)
t1 = [timeit(lambda : version_1(n), number=3) for n in order]
t2 = [timeit(lambda : version_2(n), number=3) for n in order]
t3 = [timeit(lambda : version_3(n), number=3) for n in order]
t4 = [timeit(lambda : version_4(n), number=3) for n in order]
fig, ax = plt.subplots()
for ii, t in enumerate([t1, t2, t3, t4]):
ax.plot(t, label=f"Version no. {ii+1}")
ax.set_yscale('log')
ax.set_ylabel('Execution time [s]')
ax.set_xlabel('Graph order')
ax.legend()
plt.show()Stack Overflow用户
发布于 2022-03-30 23:47:07
与使用nx.is_isomorphic比较两个图G1和G2不同,您还可以生成与G1同构的所有图,并检查G2是否在这个集合中。起初,这听起来更麻烦,但它不仅允许您检查G2是否与G1同构,还可以检查任何图是否与G1同构,而nx.is_isomorphic在比较两个图时总是从头开始。
为了让事情变得更简单,每个图都被存储成一个边列表。如果所有边的集合相同,则两个图是相同的(而不是同构的)。总是确保边的列表是一个排序的元组,这样==就可以准确地测试这个等式,并使边缘列表成为可选的。
import itertools
def all_digraphs(n):
possible_edges = [
(i, j) for i, j in itertools.product(range(n), repeat=2) if i != j
]
for edge_mask in itertools.product([True, False], repeat=len(possible_edges)):
# The result is already sorted
yield tuple(edge for include, edge in zip(edge_mask, possible_edges) if include)
def unique_digraphs(n):
already_seen = set()
for graph in all_digraphs(n):
if graph not in already_seen:
yield graph
already_seen |= {
tuple(sorted((perm[i], perm[j]) for i, j in graph))
for perm in itertools.permutations(range(n))
}与以前解决方案中的变体相比,这给出了我的机器上的下列时间:
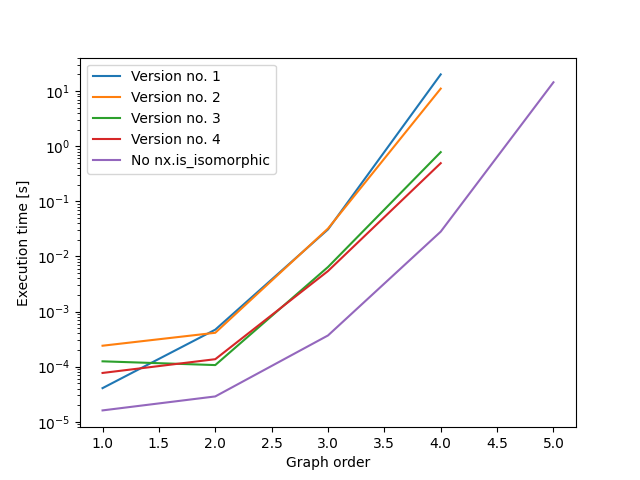
这一切看起来都很有希望,但是对于6个节点来说,我的16 for内存是不够的,Python进程已经被操作系统终止了。我相信,您可以将这段代码与为每个outdegree_sequence批量生成图结合在一起,如前面的答案所详述的那样。这将允许在每个批处理之后清空already_seen,并大大减少内存消耗。
https://stackoverflow.com/questions/71597789
复制相似问题

