如何在闪亮的应用程序中修改情节大小?
如何在闪亮的应用程序中修改情节大小?
提问于 2022-03-19 22:13:59
我创建了一个闪亮的应用程序,展示了几种情节类型。然而,地块是从顶部剪下来的,而且太宽了。我尝试修改plotOutput函数中的宽度和高度,但没有工作。
我的代码:
ui <- fluidPage(theme = shinytheme('united'),
titlePanel(title = h3("Graphs - ordered chronologically", align="center")),
selectInput("Plot",
"Choose what plots to present",
choices = list(Heatmap = "Heatmap", PCA = "PCA", VolcanoPlot = "VolcanoPlot", GSEA = 'GSEA')),
submitButton(text = "Show plots"),
verticalLayout( plotOutput(outputId = "PART.1", width = '70%'))
)这个问题在热图和火山图中最为突出。火山的地块是从山顶切下来的,热图太宽了。
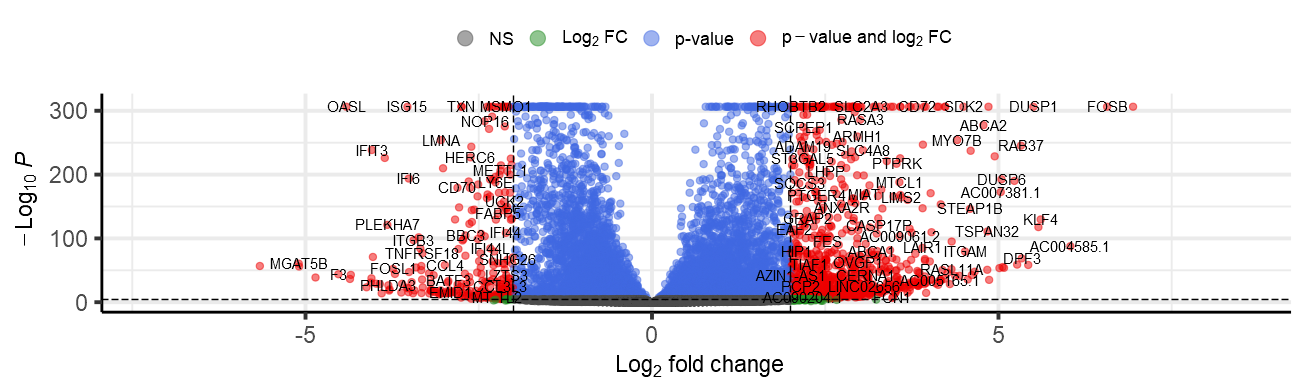
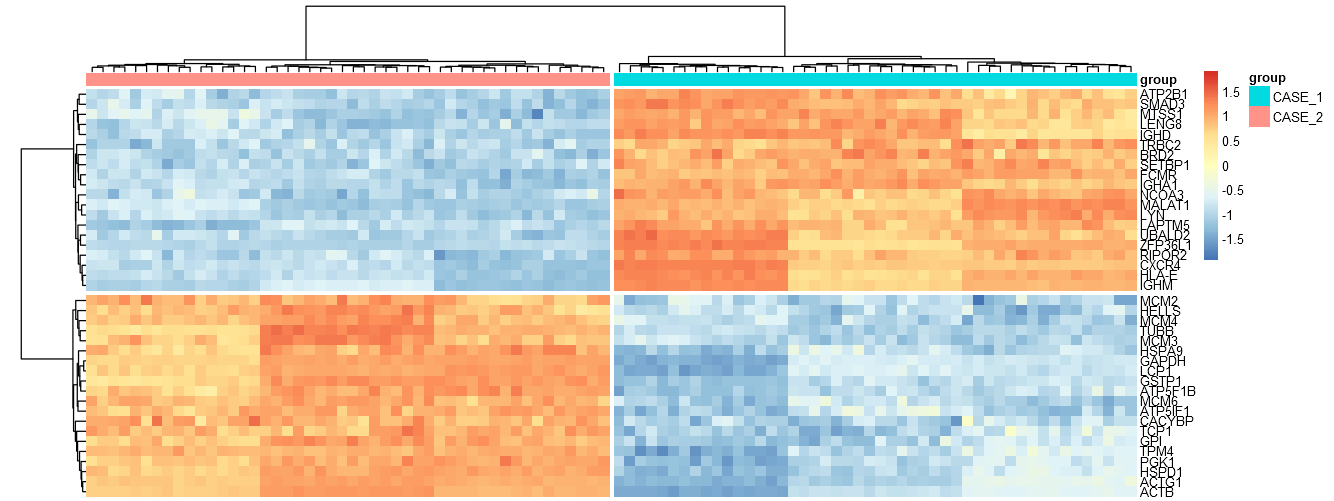
我怎么才能解决这个问题?谢谢。
编辑:
我的代码的最低版本:
library(data.table)
library(dplyr)
library(shiny)
library(shinythemes)
library(plotly)
library(compGenomRData)
library(BiocManager)
library(DESeq2)
library(org.Hs.eg.db)
library(TxDb.Hsapiens.UCSC.hg19.knownGene)
library(EnsDb.Hsapiens.v86)
library(AnnotationHub)
library(AnnotationDbi)
library(pheatmap)
library(EnhancedVolcano)
library(ggplot2)
library(FactoMineR)
library(devtools)
library(clusterProfiler)
library(ggnewscale)
library(enrichplot)
library(msigdbr)
library(readxl)
library(ExperimentHub)
library(annotate)
ui <- fluidPage(theme = shinytheme('united'),
titlePanel(title = h3("Graphs - ordered chronologically", align="center")),
selectInput("Plot",
"Choose which plots to present",
choices = list(Heatmap = "Heatmap", PCA = "PCA", VolcanoPlot = "VolcanoPlot", GSEA = 'GSEA')),
submitButton(text = "Show plots"),
verticalLayout( plotOutput(outputId = "PART.1", width = '70%'),
plotOutput(outputId = "PART.2", width = '70%'),
plotOutput(outputId = "PART.3", width = '70%'),
plotOutput(outputId = "PART.4", width = '70%'),
plotOutput(outputId = "PART.5", width = '70%'),
plotOutput(outputId = "PART.6", width = '70%'),
plotOutput(outputId = "PART.7", width = '70%'))
)
server <- function(input, output) {
# RNA-seq data
raw_counts <- frea
d('E-MTAB-7805-raw-counts.tsv', data.table = F)
metadata <- fread('E-MTAB-7805-experiment-design.tsv' ,data.table = F)
A=duplicated(raw_counts$`Gene ID`) # Check for duplicates and remove them
raw_counts = raw_counts[!A,]
A=duplicated(raw_counts$`Gene Name`) # Check for duplicates and remove them
raw_counts = raw_counts[!A,]
Hugo.Symbol <- raw_counts[,c(1:2)]
rownames(raw_counts) <- raw_counts$`Gene Name` # renaming rownames
raw_counts <- raw_counts[, -c(1:2)]
# metadata
C = duplicated(metadata$Run) # Check for duplicates and remove them
metadata = metadata[!C,]
rownames(metadata) <- metadata$Run
metadata <- metadata[,-1]
ind <- order(colnames(raw_counts), rownames(metadata))
raw_counts <- raw_counts[,ind]
# filter
target1 <- c("0 day", "1 day")
Meta_filter1 <- metadata %>% dplyr::filter(`Factor Value[time]` %in% target1)
Counts_filter1 <- raw_counts[intersect(names(raw_counts), rownames(Meta_filter1))]
rownames(Counts_filter1) <- Hugo.Symbol$`Gene Name`
# annotate
Meta_filter1$group <- plyr::mapvalues(Meta_filter1$`Factor Value[time]`, c("0 day", "1 day"),
c("CTRL", "CASE"))
ind <- order(colnames(Counts_filter1), rownames(Meta_filter1))
Counts_filter1 <- Counts_filter1[,ind]
dds <- DESeqDataSetFromMatrix(countData = Counts_filter1,
colData = Meta_filter1,
design = ~ group)
dds = DESeq(dds)
res = results(dds)
res$symbol <- rownames(res)
resOrder <- res[order(res$padj),]
# heatmap
dds.symbol = dds
rownames(dds.symbol) = mapIds(org.Hs.eg.db,
keys=rownames(dds),
column="SYMBOL",
keytype="SYMBOL",
multiVals="first")
rownames(dds.symbol)[is.na(rownames(dds.symbol))] = rownames(dds)[is.na(rownames(dds.symbol))]
rownames(dds.symbol) = make.unique(rownames(dds.symbol))
selectUp <- resOrder$symbol[resOrder$log2FoldChange>0][1:20]
selectDown <- resOrder$symbol[resOrder$log2FoldChange<0][1:20]
select = c(selectUp,selectDown)
df <- data.frame(row.names = colnames(dds.symbol),
group = colData(dds.symbol)$group)
normcounts = assay(vst(dds.symbol,blind=T, nsub = 2000))
# Functional enrichment
res = res[!is.na(res1$padj),]
mygenes <- rownames(res)
lfc = res1$log2FoldChange # get gene symbol
names(lfc) <- rownames(res)
lfc <- sort(lfc, decreasing = TRUE)
hallmarks <- msigdbr(species = "Homo sapiens", category = "H") %>%
dplyr::select(gs_name, gene_symbol)
## Output
output$PART.1 <- renderPlot({
if (input$Plot == 'Heatmap') {
pheatmap(normcounts[select,], cluster_rows=TRUE,
show_colnames = FALSE,cluster_cols=TRUE,
annotation_col=df, scale = 'row',cutree_cols = 2,cutree_rows = 2)
} else if (input$Plot == 'PCA') {
Var <- apply(normcounts, 1, var)
selectedVarGenes <- names(Var[order(Var, decreasing = T)][1:1000])
M <- t(normcounts[selectedVarGenes,])
pcaResults = prcomp(M)
qplot(pcaResults$x[,1],pcaResults$x[,2], col=dds1$group,size=2)
} else if (input$Plot == 'VolcanoPlot') {
EnhancedVolcano(resOrder,
lab = resOrder$symbol,
x = 'log2FoldChange',
y = 'padj',
labSize=4,
FCcutoff=2 )
} else {
em <- GSEA(lfc, TERM2GENE = hallmarks)
dotplot(em)
}
})
shinyApp(ui = ui, server = server)回答 1
Stack Overflow用户
发布于 2022-03-23 14:20:47
如果在调用par之前包括对pheatmap的调用,可能会有所帮助:
par(mar=c(5,4,6,2)) # bottom, left, top, right
pheatmap(
normcounts[select,], cluster_rows=TRUE,
show_colnames = FALSE, cluster_cols=TRUE,
annotation_col=df, scale = 'row',
cutree_cols = 2,cutree_rows = 2
)页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/71542515
复制相关文章
相似问题

