如何从CausalImpact包中添加图例到绘图?
我看到了关于这个的另一篇文章这里,但是我对R相对来说是比较新的,所以答案对我没有帮助。我真的很感激你能给我更多的帮助。
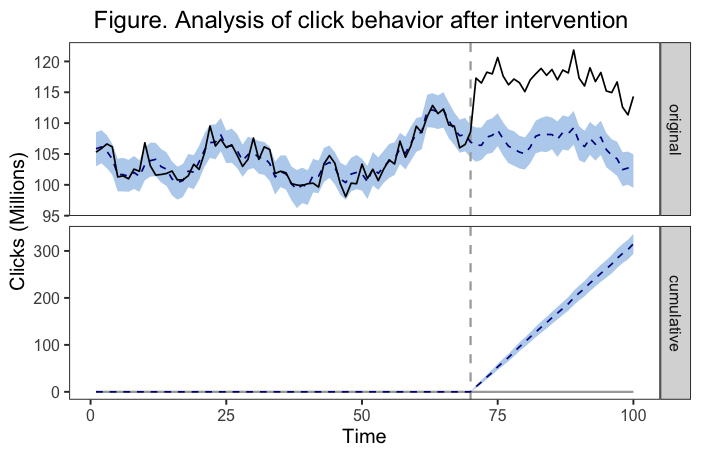
我已经用因果影响包中的命令绘制了一个图表。在包文档中,它清楚地说明这些情节是ggplot2对象,可以像其他任何类似的对象一样进行定制。我已经成功地做到了,添加标题和定制颜色。我需要添加一个图例(这是我提交给的日记所要求的)。下面是一个例子,说明我的图形现在是什么样子,以及我用来获得的代码。
library(ggplot2)
devtools::install_github("google/CausalImpact")
library(CausalImpact)
## note that I took this example code from the package documentation up until I customize the plot
#create data
set.seed(1)
x1 <- 100 + arima.sim(model = list(ar = 0.999), n = 100)
y <- 1.2 * x1 + rnorm(100)
y[71:100] <- y[71:100] + 10
data <- cbind(y, x1)
#causal impact analysis
> pre.period <- c(1, 70)
> post.period <- c(71, 100)
> impact <- CausalImpact(data, pre.period, post.period)
#graph
example<-plot(impact, c("original", "cumulative")) +
labs(
x = "Time",
y = "Clicks (Millions)",
title = "Figure. Analysis of click behavior after intervention.") +
theme(plot.title = element_text(hjust = 0.5),
plot.caption = element_text(hjust = 0),
panel.background = element_rect(fill = "transparent"), # panel bg
plot.background = element_rect(fill = "transparent", color = NA), # plot bg
panel.grid.major = element_blank(), # get rid of major grid
panel.grid.minor = element_blank()) # get rid of minor grid在我的头脑中,我想要的解决方案是有一个图板的每一个图板。第一个图例(在“原始”面板旁边)将显示一条实线表示观测数据,虚线表示估计的反事实,而彩色带表示估计反事实周围95%的CrI。第二个图例(在“累积”面板旁边)将显示虚线表示与干预相关的趋势的估计变化,而有色带再次表示估计值周围的95%的CrI。也许有一个更好的解决方案,但这正是我所想的。
下面是在绘图时运行的底层代码的一个部分:
# Initialize plot
q <- ggplot(data, aes(x = time)) + theme_bw(base_size = 15)
q <- q + xlab("") + ylab("")
if (length(metrics) > 1) {
q <- q + facet_grid(metric ~ ., scales = "free_y")
}
# Add prediction intervals
q <- q + geom_ribbon(aes(ymin = lower, ymax = upper),
data, fill = "slategray2")
# Add pre-period markers
xintercept <- CreatePeriodMarkers(impact$model$pre.period,
impact$model$post.period,
time(impact$series))
q <- q + geom_vline(xintercept = xintercept,
colour = "darkgrey", size = 0.8, linetype = "dashed")
# Add zero line to pointwise and cumulative plot
q <- q + geom_line(aes(y = baseline),
colour = "darkgrey", size = 0.8, linetype = "solid",
na.rm = TRUE)
# Add point predictions
q <- q + geom_line(aes(y = mean), data,
size = 0.6, colour = "darkblue", linetype = "dashed",
na.rm = TRUE)
# Add observed data
q <- q + geom_line(aes(y = response), size = 0.6, na.rm = TRUE)
return(q)
}在这篇老文章中,有一个答案是,我必须调整已有的功能,才能得到一个传奇,而且我还没有真正的技能去了解我需要改变或添加的东西。我原以为传说应该根据全球图形代码中的aes()位自动添加,所以我有点搞不懂为什么一开始就没有。有人能帮我吗?
回答 3
Stack Overflow用户
发布于 2022-02-12 17:53:50
以下是早期解决方案的更新/编辑版本,以便将美学合并到一个图例中。要求是将线型和填充(色带颜色)合并成一个图例。
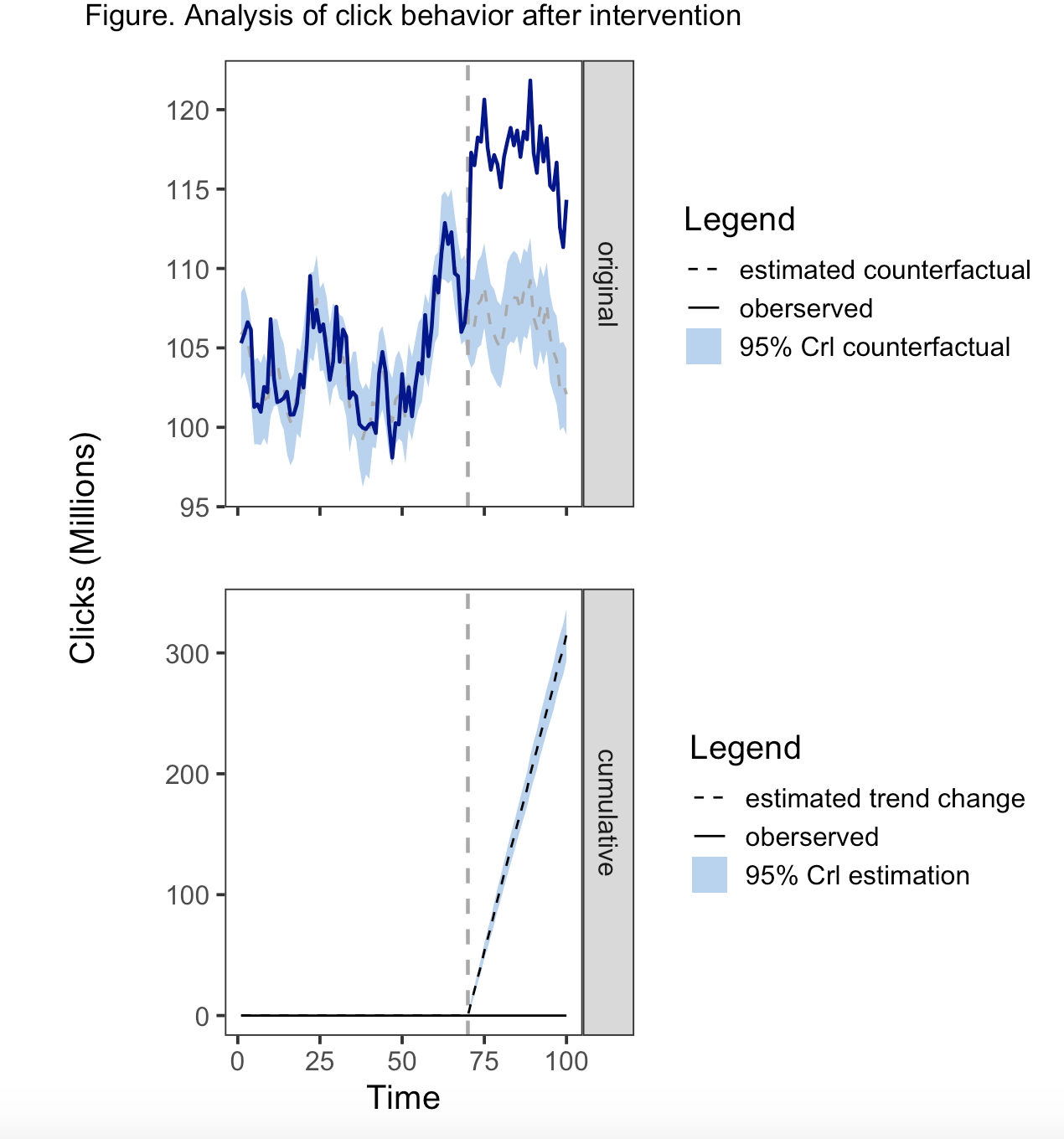
为了合并传说,同样的美学必须在地理上使用,尺度必须考虑不同的变量,具有相同的名称和相同的标签。因此,geom_ribbon()需要在aes()和geom_line()中都有一个行类型,而geom_line()需要填充aes()和linetype。将线型添加到geom_ribbon()中的一个副作用是,在带的两边都有一条线。另一方面,fill不适用于geom_line,因此您只会收到一条警告信息,即填充美学将被忽略。
解决这一问题的方法是对scale_linetype_manual()中的相关值应用“空白”行类型。类似地,我们在scale_fill_manual()中使用“透明”来避免将颜色应用于比例的其他元素。
在研究这个之前,我没有意识到的是,为多个变量的价值创造一个传奇是可能的。这些值只需在比例中适当地映射即可。所以我真的学到了一些新东西--把它组合在一起。
CreateImpactPlot <- function(impact, metrics = c("original", "cumulative")) {
# Creates a plot of observed data and counterfactual predictions.
#
# Args:
# impact: \code{CausalImpact} results object returned by
# \code{CausalImpact()}.
# metrics: Which metrics to include in the plot. Can be any combination of
# "original", "pointwise", and "cumulative".
#
# Returns:
# A ggplot2 object that can be plotted using plot().
# Create data frame of: time, response, mean, lower, upper, metric
data <- CreateDataFrameForPlot(impact)
# Select metrics to display (and their order)
assert_that(is.vector(metrics))
metrics <- match.arg(metrics, several.ok = TRUE)
data <- data[data$metric %in% metrics, , drop = FALSE]
data$metric <- factor(data$metric, metrics)
# Make data longer
data_long <- data %>%
tidyr::pivot_longer(cols = c("baseline", "mean", "response"), names_to = "variable",
values_to = "value", values_drop_na = TRUE)
# Initialize plot
q1 <- ggplot(data, aes(x = time)) + theme_bw(base_size = 15)
q1 <- q1 + xlab("") + ylab("")
q3 <- ggplot(data %>%
filter(metric == "cumulative") %>%
mutate(metric = factor(metric, levels = c("cumulative"))), aes(x = time)) + theme_bw(base_size = 15)
q3 <- q3 + xlab("") + ylab("")
# Add prediction intervals
q1 <- q1 + geom_ribbon(data = data %>%
filter(metric == "original") %>%
mutate(metric = factor(metric, levels = c("original"))), aes(x = time, ymin = lower, ymax = upper, fill = metric,
linetype = metric))
q3 <- q3 + geom_ribbon(data = data %>%
filter(metric == "cumulative") %>%
mutate(metric = factor(metric, levels = c("cumulative"))), aes(x = time, ymin = lower, ymax = upper, fill = metric))
# Add pre-period markers
xintercept <- CreatePeriodMarkers(impact$model$pre.period,
impact$model$post.period,
time(impact$series))
q1 <- q1 + geom_vline(xintercept = xintercept,
colour = "darkgrey", size = 0.8, linetype = "dashed")
q3 <- q3 + geom_vline(xintercept = xintercept,
colour = "darkgrey", size = 0.8, linetype = "dashed")
# Add zero line to cumulative plot
# Add point predictions
# Add observed data
q1 <- q1 + geom_line(data = data_long %>% dplyr::filter(metric == "original"),
aes(x = time, y = value, linetype = variable, group = variable,
size = variable, fill = variable, color = variable),
na.rm = TRUE)+
scale_linetype_manual(name = "Legend", labels = c("mean"= "estimated counterfactual", "response" = "oberserved", "original" = "95% Crl counterfactual"),
values = c("dashed", "solid", "blank"), limits = c("mean", "response","original")) +
scale_fill_manual(name = "Legend", labels = c("mean"= "estimated counterfactual", "response" = "oberserved", "original" = "95% Crl counterfactual"),
values = c("transparent", "transparent","slategray2"), limits = c("mean", "response","original")) + #limits controls the order in the legend
scale_size_manual(values = c(0.6, 0.8, 0.5)) +
scale_color_manual(values = c("darkgray", "darkblue")) +
theme(legend.position = "right", axis.text.x = element_blank(), axis.title.y = element_blank()) +
guides(size = "none", color = "none")+
facet_wrap(~metric[1], strip.position = "right", drop = TRUE) #use facet_wrap to generate the stip
q3 <- q3 + geom_line(data = data_long %>% dplyr::filter(metric == "cumulative"),
aes(x = time, y = value, linetype = variable, group = variable,
fill = variable),
na.rm = TRUE) +
scale_linetype_manual(name = "Legend", labels = c("mean"= "estimated trend change", "baseline" = "oberserved", "cumulative" = "95% Crl estimation"),
values = c("dashed", "solid", "blank"), limits = c("mean", "baseline","cumulative")) +
scale_fill_manual(name = "Legend", labels = c("mean"= "estimated trend change", "baseline" = "oberserved", "cumulative" = "95% Crl estimation"),
values = c("transparent", "transparent","slategray2"), limits = c("mean", "baseline","cumulative")) + #limits controls the order in the legend
theme(legend.position = "right", axis.title.y = element_blank())+
labs(x = "Time") +
facet_wrap(~metric, strip.position = "right", drop = TRUE) #use facet_wrap to generate the stip
g1 <- grid::textGrob("Clicks (Millions)", rot = 90, gp=gpar(fontsize = 15), x= 0.85)
wrap_elements(g1) | (q1/q3)
patchwork <- wrap_elements(g1) | (q1/q3)
q <- patchwork
return(q)
}
# To run the function
plot(impact, c("original", "cumulative")) +
plot_annotation(title = "Figure. Analysis of click behavior after intervention"
, theme = theme(plot.title = element_text(hjust = 0.5))) &
theme(
panel.background = element_rect(fill = "transparent"), # panel bg
plot.background = element_rect(fill = "transparent", color = NA), # plot bg
panel.grid.major = element_blank(), # get rid of major grid
panel.grid.minor = element_blank())Stack Overflow用户
发布于 2022-02-12 01:55:12
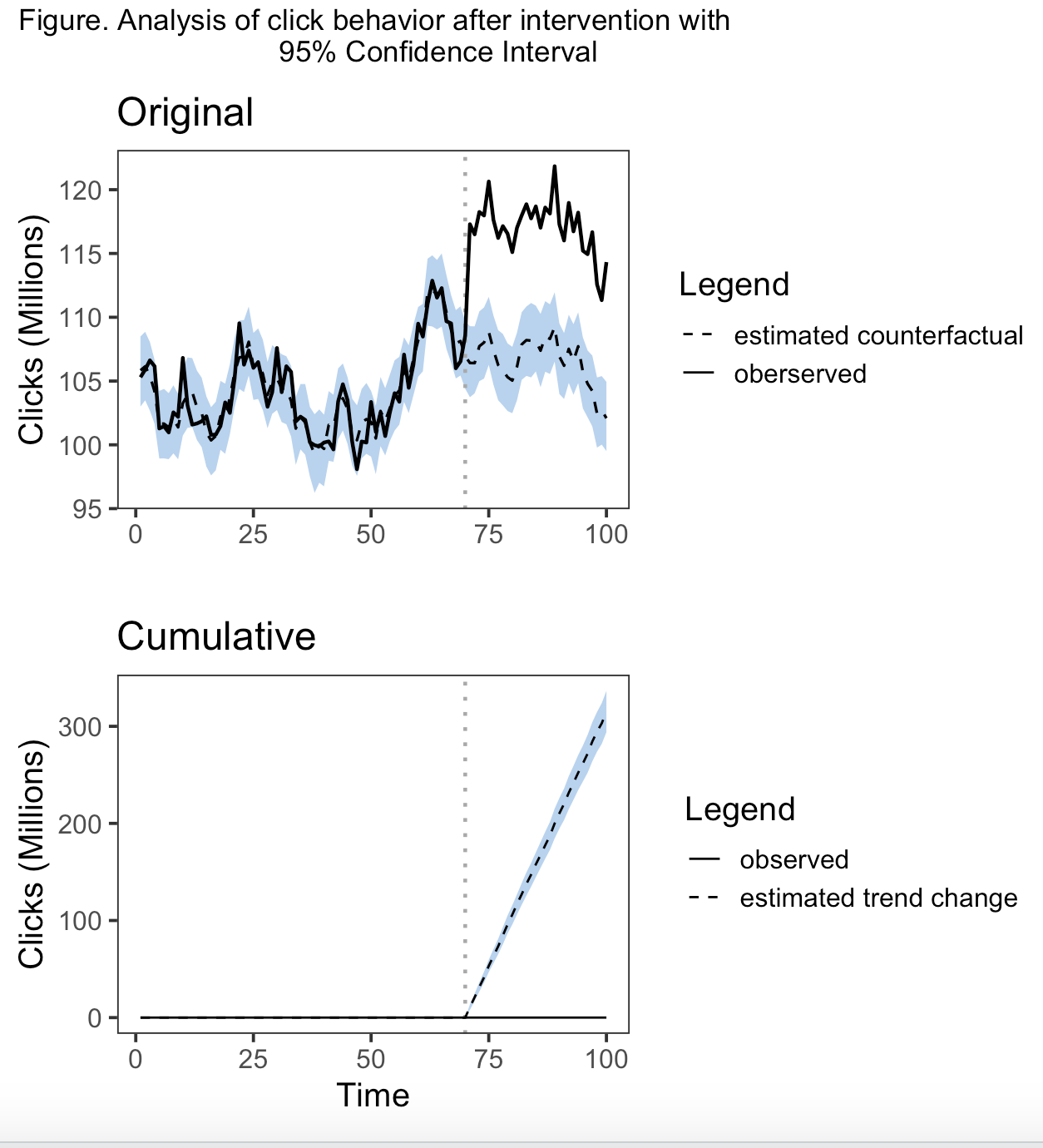
我重写了情节函数。我没有使用facet_wrap(),而是用自己的传说创建了单独的情节,并使用拼凑的方法将它们组合成一个单独的情节。为了运行这一点,您需要存储所有源代码,包括impact_analysis.R、impact_misc.R、impact_model.R、impact_inference.R和impact_plot.R,但我重新创建的CreateImpactPlot函数除外。因此,相反,运行我有以下内容。您还需要加载ggplot2、tidyr、dplyr和修补程序。这将只运行原始和累积的指标。虽然我在某种程度上修改了Pointwise,但我不想这样做,因为我没有一个可复制的例子。我将您的主题首选项直接输入到函数中的代码中。你应该能够在闲暇的时候识别和改变这些元素。要明确的是,这些情节是q1 =原始的,q2 =点态,q3 =累积的。我不知道如何将信任带带入传奇,因为它不是aes()的一部分。可能会从零开始创建一个grob。我只是在标题中引用了它,如果不适合你,你可以修改它。希望这能帮上忙。
"cumulative")) {
# Creates a plot of observed data and counterfactual predictions.
#
# Args:
# impact: \code{CausalImpact} results object returned by
# \code{CausalImpact()}.
# metrics: Which metrics to include in the plot. Can be any combination of
# "original", "pointwise", and "cumulative".
#
# Returns:
# A ggplot2 object that can be plotted using plot().
# Create data frame of: time, response, mean, lower, upper, metric
data <- CreateDataFrameForPlot(impact)
# Select metrics to display (and their order)
assert_that(is.vector(metrics))
metrics <- match.arg(metrics, several.ok = TRUE)
data <- data[data$metric %in% metrics, , drop = FALSE]
data$metric <- factor(data$metric, metrics)
# Initialize plot
#q <- ggplot(data, aes(x = time)) + theme_bw(base_size = 15)
#q <- q + xlab("") + ylab("")
#if (length(metrics) > 1) {
# q <- q + facet_grid(metric ~ ., scales = "free_y")
#}
q1 <- ggplot(data, aes(x = time)) + theme_bw(base_size = 15)
q1 <- q1 + xlab("") + ylab("")
q2 <- ggplot(data, aes(x = time)) + theme_bw(base_size = 15)
q2 <- q2 + xlab("") + ylab("")
q3 <- ggplot(data, aes(x = time)) + theme_bw(base_size = 15)
q3 <- q3 + xlab("") + ylab("")
# Add prediction intervals
#q <- q + geom_ribbon(aes(ymin = lower, ymax = upper),
# data, fill = "slategray2")
q1 <- q1 + geom_ribbon(data = data %>% dplyr::filter(metric == "original"), aes(x = time, ymin = lower, ymax = upper),
fill = "slategray2")
q2 <- q2 + geom_ribbon(data = data %>% dplyr::filter(metric == "pointwise"), aes(x = time, ymin = lower, ymax = upper),
fill = "slategray2")
q3 <- q3 + geom_ribbon(data = data %>% dplyr::filter(metric == "cumulative"), aes(x = time, ymin = lower, ymax = upper),
fill = "slategray2")
# Add pre-period markers
xintercept <- CreatePeriodMarkers(impact$model$pre.period,
impact$model$post.period,
time(impact$series))
#q <- q + geom_vline(xintercept = xintercept,
# colour = "darkgrey", size = 0.8, linetype = "dashed")
q1 <- q1 + geom_vline(xintercept = xintercept,
colour = "darkgrey", size = 0.8, linetype = "dotted")
q2 <- q2 + geom_vline(xintercept = xintercept,
colour = "darkgrey", size = 0.8, linetype = "dotted")
q3 <- q3 + geom_vline(xintercept = xintercept,
colour = "darkgrey", size = 0.8, linetype = "dotted")
data_long <- data %>%
tidyr::pivot_longer(cols = c("baseline", "mean", "response"), names_to = "variable",
values_to = "value", values_drop_na = TRUE)
# Add zero line to pointwise and cumulative plot
#q <- q + geom_line(aes(y = baseline),
# colour = "darkgrey", size = 0.8, linetype = "solid",
# na.rm = TRUE)
q1 <- q1 + geom_line(data = data_long %>% dplyr::filter(metric == "original"),
aes(x = time, y = value, linetype = variable, group = variable,
size = variable),
na.rm = TRUE)+
scale_linetype_manual(guide = "Legend", labels = c("estimated counterfactual", "oberserved"),
values = c("dashed", "solid")) +
scale_size_manual(values = c(0.6, 0.8)) +
scale_color_manual(values = c("darkblue", "darkgrey")) +
theme(legend.position = "right") +
guides(linetype = guide_legend("Legend", nrow=2), size = "none", color = "none")+
labs(title = "Original", y = "Clicks (Millions)") +
theme(
panel.background = element_rect(fill = "transparent"), # panel bg
plot.background = element_rect(fill = "transparent", color = NA), # plot bg
panel.grid.major = element_blank(), # get rid of major grid
panel.grid.minor = element_blank())
#q2 <- q2 + geom_line(data = data_long %>% dplyr::filter(metric == "pointwise"),
# aes(x = time, y = value, linetype = Line, group = Line),
# na.rm = TRUE) +
# scale_linetype_manual(title = "Legend", labels = c("estimated counterfactual", "observed"),
# values = c("dashed", "solid")) +
# scale_size_manual(values = c(0.6, 0.8)) +
# scale_color_manual(values = c("darkblue", "darkgrey")) +
# theme(legend.position = "right") +
# guides(linetype = guide_legend("Legend", nrow=2), size = "none", color = "none")+
# labs(title = "Pointwise", y = "Clicks (Millions)")
q3 <- q3 + geom_line(data = data_long %>% dplyr::filter(metric == "cumulative"),
aes(x = time, y = value, linetype = variable, group = variable),
na.rm = TRUE) +
scale_linetype_manual(labels = c("observed", "estimated trend change"),
values = c("solid", "dashed")) +
theme(legend.position = "right")+
guides(linetype = guide_legend("Legend", nrow=2))+
labs(title = "Cumulative",x = "Time", y = "Clicks (Millions)")+
theme(
panel.background = element_rect(fill = "transparent"), # panel bg
plot.background = element_rect(fill = "transparent", color = NA), # plot bg
panel.grid.major = element_blank(), # get rid of major grid
panel.grid.minor = element_blank())
patchwork <- q1 / q3
q <- patchwork + plot_annotation(title = "Figure. Analysis of click behavior after intervention with
95% Confidence Interval")
# Add point predictions
#q <- q + geom_line(aes(y = mean), data,
# size = 0.6, colour = "darkblue", linetype = "dashed",
# na.rm = TRUE)
# Add observed data
#q <- q + geom_line(aes(y = response), size = 0.6, na.rm = TRUE)
return(q)
}
plot(impact, c("original", "cumulative"))
Stack Overflow用户
发布于 2022-02-13 02:25:48
下面是对CreateImpactPlot()函数的重新构建,它将适用于所有三个指标。传说是可以修改的。我引入了更多的颜色和线型,以便传奇可以适用于所有方面。
基本情况如下:
plot(impact)
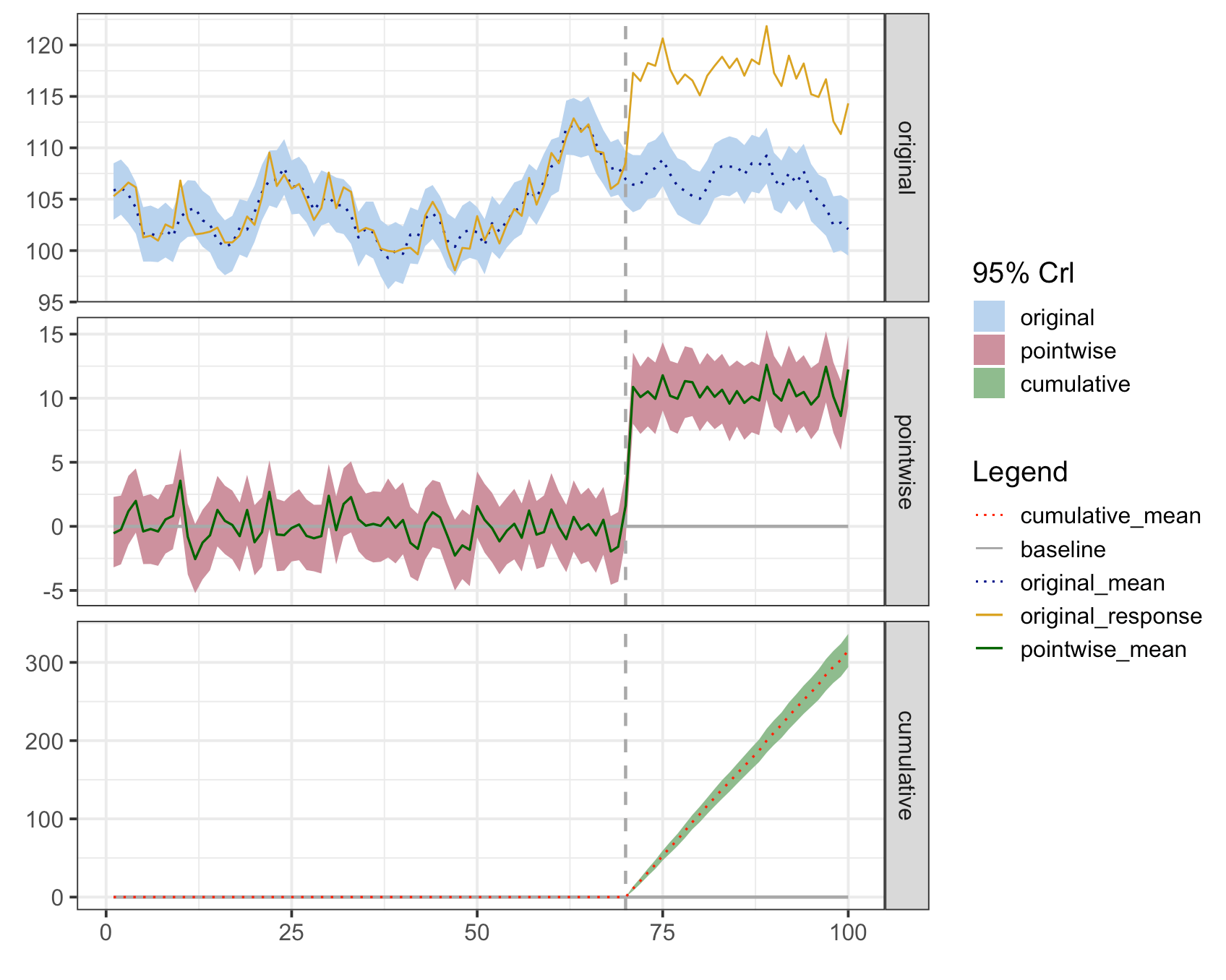
您将注意到,图例中的缎带和线条的标签引用了度量标准。这些是占位符标签,然后可以修改。
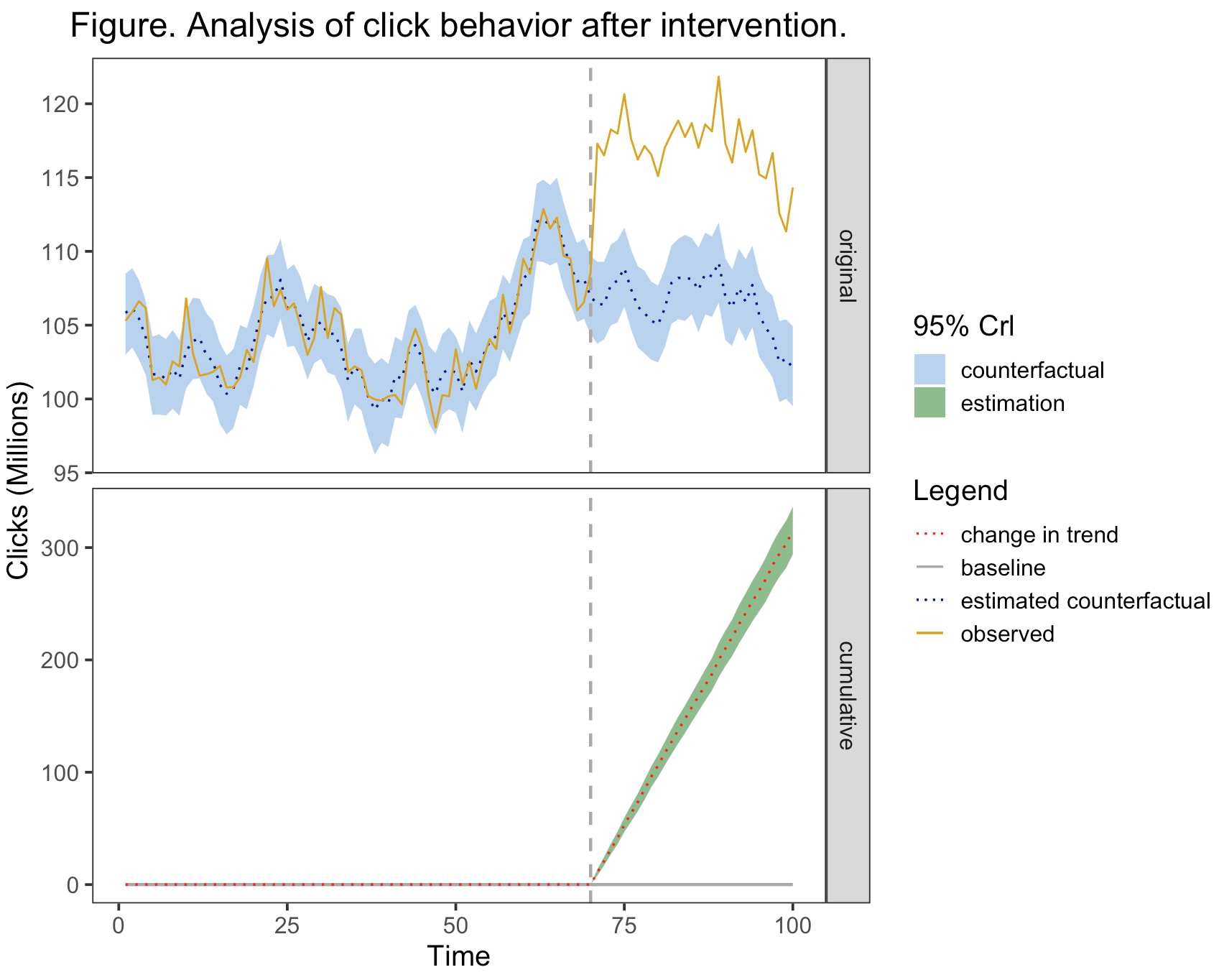
line_labels <- c("cumulative_mean" = "change in trend", "baseline" = "baseline", "original_mean" =
"estimated counterfactual", "original_response" = "observed")
plot(impact, c("original", "cumulative")) +
labs(
x = "Time",
y = "Clicks (Millions)",
title = "Figure. Analysis of click behavior after intervention.") +
theme(plot.title = element_text(hjust = 0.5),
plot.caption = element_text(hjust = 0),
panel.background = element_rect(fill = "transparent"), # panel bg
plot.background = element_rect(fill = "transparent", color = NA), # plot bg
panel.grid.major = element_blank(), # get rid of major grid
panel.grid.minor = element_blank()) + # get rid of minor grid
scale_fill_manual(name = "95% Crl", values = c("original" = "slategray2", "cumulative" = "darkseagreen"),
labels = c("original" = "counterfactual", "cumulative" = "estimation")) +
scale_linetype_manual(name = "Legend", labels = line_labels,
values = c("cumulative_mean" = "dotted", "baseline" = "solid", "original_mean" =
"dotted", "original_response" = "solid")) +
scale_color_manual(name = "Legend", labels = line_labels,
values = c("cumulative_mean" = "red", "baseline" = "darkgrey", "original_mean"= "darkblue", "original_response" = "goldenrod"))向量"line_labels“是定义要在”图解“中显示的文本的地方。您将注意到,我删除了点相关值,因为我将点态度量从绘图中排除在外。scale_linetype_manual和scale_color_manual必须保持名称和标签的同步,这样才能有一个合并的传奇,否则就会有两个不同的传说。scale_fill_manual是给缎带的。对于这些比例,您可以根据需要更改名称、标签和值。您可以从函数中复制代码,修改它,并将其添加到上面所示的绘图调用中。
这是修改后的函数的代码。在本例中,应该运行所有内容,并从CausalImpact包中生成“影响”。然后,所有包代码都需要加载到内存中,包括impact_analysis.R、impact_misc.R、impact_model.R、impact_inference.R和impact_plot.R,然后加载下面的代码。
CreateImpactPlot2 <- function(impact, metrics = c("original", "pointwise","cumulative")) {
# Creates a plot of observed data and counterfactual predictions.
#
# Args:
# impact: \code{CausalImpact} results object returned by
# \code{CausalImpact()}.
# metrics: Which metrics to include in the plot. Can be any combination of
# "original", "pointwise", and "cumulative".
#
# Returns:
# A ggplot2 object that can be plotted using plot().
# Create data frame of: time, response, mean, lower, upper, metric
data <- CreateDataFrameForPlot(impact)
# Select metrics to display (and their order)
assert_that(is.vector(metrics))
metrics <- match.arg(metrics, several.ok = TRUE)
data <- data[data$metric %in% metrics, , drop = FALSE]
data$metric <- factor(data$metric, metrics)
data_long <- data %>%
tidyr::pivot_longer(cols = c("baseline", "mean", "response"), names_to = "variable",
values_to = "value", values_drop_na = TRUE) %>%
mutate(variable2 = factor(ifelse(variable == "baseline", variable, paste0(metric,"_", variable))),
variable = factor(variable))
# Initialize plot
q <- ggplot(data, aes(x = time)) + theme_bw(base_size = 15)
q <- q + xlab("") + ylab("")
if (length(metrics) > 1) {
q <- q + facet_grid(metric ~ ., scales = "free_y")
}
#Add prediction intervals
q <- q + geom_ribbon(aes(x = time, ymin = lower, ymax = upper, fill = metric), data_long)
# Add pre-period markers
xintercept <- CreatePeriodMarkers(impact$model$pre.period,
impact$model$post.period,
time(impact$series))
q <- q + geom_vline(xintercept = xintercept,
colour = "darkgrey", size = 0.8, linetype = "dashed")
# Add zero line to pointwise and cumulative plot
q <- q + geom_line(data = data_long %>% dplyr::filter(variable == "baseline"),
aes(x = time, y = value, linetype = variable2, group = variable2, size = variable2, color = variable2),
na.rm = TRUE)
# Add point predictions
q <- q + geom_line(data = data_long %>% dplyr::filter(variable == "mean"),
aes(x = time, y = value, linetype = variable2, group = variable2, size = variable2, color = variable2),
na.rm = TRUE)
# Add observed data
q <- q + geom_line(data = data_long %>% dplyr::filter(variable == "response"),
aes(x = time, y = value, linetype = variable2, group = variable2, size = variable2, color = variable2),
na.rm = TRUE)
#Add scales
line_labels <- c("cumulative_mean" = "cumulative_mean", "baseline" = "baseline", "original_mean" =
"original_mean", "original_response" = "original_response", "pointwise_mean"=
"pointwise_mean")
q <- q + scale_linetype_manual(name = "Legend", labels = line_labels,
values = c("cumulative_mean" = "dotted", "baseline" = "solid", "original_mean" =
"dotted", "original_response" = "solid", "pointwise_mean"=
"solid")) +
scale_size_manual(values = c("cumulative_mean" = 0.6, "baseline" = 0.8, "original_mean"= 0.6, "original_response" = 0.5,
"pointwise_mean"= 0.6)) +
scale_color_manual(name = "Legend", labels = line_labels,
values = c("cumulative_mean" = "red", "baseline" = "darkgrey", "original_mean"= "darkblue", "original_response" = "goldenrod",
"pointwise_mean"= "darkgreen")) +
scale_fill_manual(name = "95% Crl", values = c("original" = "slategray2", "pointwise" = "pink3", "cumulative" = "darkseagreen"),
labels = c("original" = "original", "pointwise" = "pointwise", "cumulative" = "cumulative")) +
guides(size = "none")
return(q)
}
plot.CausalImpact <- function(x, ...) {
# Creates a plot of observed data and counterfactual predictions.
#
# Args:
# x: A \code{CausalImpact} results object, as returned by
# \code{CausalImpact()}.
# ...: Can be used to specify \code{metrics}, which determines which panels
# to include in the plot. The argument \code{metrics} can be any
# combination of "original", "pointwise", "cumulative". Partial matches
# are allowed.
#
# Returns:
# A ggplot2 object that can be plotted using plot().
#
# Examples:
# \dontrun{
# impact <- CausalImpact(...)
#
# # Default plot:
# plot(impact)
#
# # Customized plot:
# impact.plot <- plot(impact) + ylab("Sales")
# plot(impact.plot)
# }
return(CreateImpactPlot2(x, ...))
}https://stackoverflow.com/questions/71073879
复制相似问题

