如何优化熊猫的巢式循环?
编辑27/01/2022
我将测试@hpchavaz的解决方案
缓慢问题的一部分来自调用循环期间的函数。其中2个函数加载csv文件(pd.read_csv)。我通过只加载所需的列(而不是所有文件)进行优化。
之后,我试着避免循环:
- 使用pd.melt在行中“转置”列
- 使用pd.melt
我将执行时间除以5 (227分钟-> 45分钟)
pd.set_option("display.max_rows", None, "display.max_columns", None)
# pd.set_option('max_colwidth', None)
# page 332 of SDTMIG3.3 Guidelines
# Finding About
domain = 'FA'
print(f'Domain {domain}')
print(datetime.datetime.now().strftime('%Y-%m-%d - %H:%M:%S'))
start = datetime.datetime.now()
# Données sources
df_randomisation = pd.read_csv(f'./csv/source/randomisation.csv',delimiter=",")
df_randomisation = df_randomisation.loc[:, df_randomisation.columns!='redcap_repeat_instance'] # exclude column redcap_repeat_instance as randomisation form are unique
df_randomisation = df_randomisation.loc[:, df_randomisation.columns!='redcap_event_name'] # exclude column redcap_repeat_name as randomisation form are unique
df_randomisation = df_randomisation.loc[:, df_randomisation.columns!='ID'] # exclude column redcap_repeat_name as randomisation form are unique
df_clinical_exam = pd.read_csv(f'./csv/source/examen_clinique_symptomes.csv',delimiter=",")
df = pd.merge(df_randomisation,df_clinical_exam, left_on='pat_ide', right_on='pat_ide', how='outer').sort_values(by=['pat_ide','sym_dat'])
# filtre sur un patient pour les tests
# df = df[(df['pat_ide'] == 'BFBO001')]
# print(df)
# Transformation des données sources : évènements en colonnes => évènements en lignes
df1 = pd.melt(df, id_vars=['pat_ide','sym_dat','redcap_event_name'], value_vars=['sym_tou', 'sym_gra','sym_ete','sym_fie','sym_fat','sym_dys','sym_mya','sym_cep','sym_gor','sym_cut','sym_ore','sym_nez','sym_odo','sym_gou','sym_ver','sym_vis','sym_aud','sym_tho','sym_pal','sym_nau','sym_vom','sym_dia','sym_abd','sym_mal','sym_con'],var_name='symptome', value_name='intensite')
df2 = pd.melt(df, id_vars=['pat_ide','sym_dat'], value_vars=['sym_tou_del','sym_gra_del','sym_ete_del','sym_fie_del','sym_fat_del','sym_dys_del','sym_mya_del','sym_cep_del','sym_gor_del','sym_cut_del','sym_ore_del','sym_nez_del','sym_odo_del','sym_gou_del','sym_ver_del','sym_vis_del','sym_aud_del','sym_tho_del','sym_pal_del','sym_nau_del','sym_vom_del','sym_dia_del','sym_abd_del','sym_mal_del','sym_con_del'],var_name='symptome', value_name='delai')
df2['symptome'] = df2.apply(lambda row: row['symptome'][:-4],axis=1)
# print(df1)
# print(df2)
# list of 25 monitored clinical signs
signs = {
'sym_tou' : 'Dry cough',
'sym_gra' : 'Fatty cough',
'sym_ete' : 'Sneezing',
'sym_fie' : 'Fever',
'sym_fat' : 'Tiredness',
'sym_dys' : 'Dyspnea',
'sym_mya' : 'Muscle aches (Myalgia)',
'sym_cep' : 'Headache',
'sym_gor' : 'Sore throat',
'sym_cut' : 'Skin rash',
'sym_ore' : 'Earache',
'sym_nez' : 'Runny nose (Rhinorrhea)',
'sym_odo' : 'Loss of smell (Anosmia)',
'sym_gou' : 'Loss of taste (Ageusia)',
'sym_ver' : 'Feeling dizzy',
'sym_vis' : 'Acute vision impairment',
'sym_aud' : 'Acute hearing impairment',
'sym_tho' : 'Chest pain',
'sym_pal' : 'Palpitations',
'sym_nau' : 'Nausea',
'sym_vom' : 'Vomiting',
'sym_dia' : 'Diarrhea',
'sym_abd' : 'Abdominal pain',
'sym_mal' : 'Inexplicated tiredness',
'sym_con' : 'Conjunctivitis'
}
dataframe = pd.DataFrame()
dataframe = pd.merge(df1,df2, left_on=['pat_ide','sym_dat','symptome'], right_on=['pat_ide','sym_dat','symptome'], how='outer').fillna('NULL')
dataframe['symptome'] = dataframe.apply(lambda row: signs[row['symptome']],axis=1)
dataframe = dataframe.sort_values(by=['pat_ide','sym_dat'])
# print(dataframe)
# print(len(dataframe.index))
# FA
sdtm_variables = ['STUDYID','DOMAIN','USUBJID','FASEQ','FATESTCD','FATEST','FAOBJ','FACAT','FAORRES','FAORRESU','FASTRESC','FASTRESN','FASTAT','VISITNUM','VISIT','VISITDY','EPOCH','FADTC','FADY','SYM_DAT','DEL','pat_ide']
df_fa = pd.DataFrame(columns = sdtm_variables)
df_fa['pat_ide'] = dataframe.apply(lambda row: row['pat_ide'],axis=1)
df_fa['STUDYID'] = dataframe.apply(lambda row: '01-COV',axis=1)
df_fa['DOMAIN'] = dataframe.apply(lambda row: 'FA',axis=1)
df_fa['USUBJID'] = dataframe.apply(lambda row: Patient_icon_format(row['pat_ide']),axis=1) # index of df
df_fa['FASEQ'] = dataframe.apply(lambda row: 1,axis=1) # pd.RangeIndex(stop=dataframe.shape[0]) + 1
df_fa['FATESTCD'] = dataframe.apply(lambda row:'OCCUR',axis=1)
df_fa['FATEST'] = dataframe.apply(lambda row:'Occurrence Indicator',axis=1)
df_fa['FAOBJ'] = dataframe.apply(lambda row: row['symptome'],axis=1)
df_fa['FACAT'] = dataframe.apply(lambda row: 'COVID-19 SYMPTOMS',axis=1)
df_fa['FAORRES'] = dataframe.apply(lambda row: Clinical_result(row['intensite']),axis=1)
df_fa['FAORRESU'] = dataframe.apply(lambda row: '',axis=1)
df_fa['FASTRESC'] = dataframe.apply(lambda row: Clinical_standard_result(row['intensite']),axis=1)
df_fa['FASTRESN'] = dataframe.apply(lambda row: np.nan,axis=1)
df_fa['FASTAT'] = dataframe.apply(lambda row: 'NOT DONE' if Clinical_result(row['intensite']) == '' else '',axis=1)
df_fa['VISITNUM'] = dataframe.apply(lambda row: str(Visit_numbering(row['pat_ide'],row['sym_dat'],row['redcap_event_name'])),axis=1)
df_fa['VISIT'] = dataframe.apply(lambda row: Visit_name(row['pat_ide'],row['sym_dat'],row['redcap_event_name']),axis=1)
df_fa['VISITDY'] = dataframe.apply(lambda row: '-1' if Visit_numbering(row['pat_ide'],row['sym_dat'],row['redcap_event_name']) == 1 else str(round((Visit_numbering(row['pat_ide'],row['sym_dat'],row['redcap_event_name']) - 1))),axis=1)
df_fa['EPOCH'] = dataframe.apply(lambda row: Get_epoch(row['pat_ide'],row['sym_dat'],row['redcap_event_name']),axis=1)
df_fa['FADTC'] = dataframe.apply(lambda row: pd.to_datetime(row['sym_dat']).strftime('%Y-%m-%dT%H:%M:%S') if row['sym_dat'] != 'NULL' else None,axis=1)
df_fa['FADY'] = dataframe.apply(lambda row: Number_of_day_since_the_subject_start_the_study(dataframe,row['sym_dat'],Patient_study_start_date(row['pat_ide'])),axis=1)
df_fa['SYM_DAT'] = dataframe.apply(lambda row: row['sym_dat'],axis=1)
df_fa['DEL'] = dataframe.apply(lambda row: row['delai'],axis=1)
# add line numbers of event (based on pat_ide and date of event) => use of redacp_repeat_instance variable of the event form
df_fa = df_fa.sort_values(by=['pat_ide','FADTC'])
df_fa = Sequence(df_fa,'FASEQ','FADTC')
df_fa = df_fa.sort_values(by=['pat_ide','FADTC'])
print(df_fa)
# SUPPFA
# creation of empty dataframes
# 2 dataframes because 2 'SUPP' variables are necessary for SUPPFA (FASTDTC et FAPRESP)
df_suppfa1 = pd.DataFrame(columns = ['STUDYID','RDOMAIN','USUBJID','IDVAR','IDVARVAL','QNAM','QLABEL','QVAL','QORIG','QEVAL','pat_ide'])
df_suppfa2 = pd.DataFrame(columns = ['STUDYID','RDOMAIN','USUBJID','IDVAR','IDVARVAL','QNAM','QLABEL','QVAL','QORIG','QEVAL','pat_ide'])
df_suppfa1['pat_ide'] = df_fa.apply(lambda row: row['pat_ide'],axis=1)
df_suppfa1['STUDYID'] = '01-COV'
df_suppfa1['RDOMAIN'] = 'FA'
df_suppfa1['USUBJID'] = df_fa.apply(lambda row: row['USUBJID'],axis=1)
df_suppfa1['IDVAR'] = 'FASEQ'
df_suppfa1['IDVARVAL'] = df_fa.apply(lambda row: row['FASEQ'],axis=1)
df_suppfa1['QNAM'] = 'FASTDTC'
df_suppfa1['QLABEL'] = 'COVID-19 Symptom Start Date'
df_suppfa1['QVAL'] = df_fa.apply(lambda row: (pd.to_datetime(row['SYM_DAT']) + timedelta(days = - int(row['DEL']))).strftime('%Y-%m-%d') if row['DEL'] != 'NULL' else row['SYM_DAT'],axis=1)
df_suppfa1['QORIG'] = 'CRF'
df_suppfa1['QEVAL'] = ''
df_suppfa2['pat_ide'] = df_fa.apply(lambda row: row['pat_ide'],axis=1)
df_suppfa2['STUDYID'] = '01-COV'
df_suppfa2['RDOMAIN'] = 'FA'
df_suppfa2['USUBJID'] = df_fa.apply(lambda row: row['USUBJID'],axis=1)
df_suppfa2['IDVAR'] = 'FASEQ'
df_suppfa2['IDVARVAL'] = df_fa.apply(lambda row: row['FASEQ'],axis=1)
df_suppfa2['QNAM'] = 'FAPRESP'
df_suppfa2['QLABEL'] = 'Prespecified'
df_suppfa2['QVAL'] = 'Y'
df_suppfa2['QORIG'] = 'CRF'
df_suppfa2['QEVAL'] = ''
dataframes = [df_suppfa1,df_suppfa2]
df_suppfa = pd.concat(dataframes)
df_suppfa = df_suppfa.drop(['pat_ide',], axis=1, errors='ignore')
# print(df_suppfa)
#-----------------------------------------------------------------------------------------------------------------------
sdtm_variables.remove('pat_ide')
sdtm_variables.remove('SYM_DAT')
sdtm_variables.remove('DEL')
# print(sdtm_variables)
df_fa = df_fa[sdtm_variables]
df_fa = df_fa.replace('NULL','')
# 3. Export data to csv
df_fa.to_csv(f'./xpt/{domain.lower()}_refactored.csv',index=False)
df_suppfa.to_csv(f'./xpt/supp{domain.lower()}_refactored.csv',index=False)
print(datetime.datetime.now().strftime('%Y-%m-%d - %H:%M:%S'))
print('Execution time',datetime.datetime.now() - start)编辑
正常情况下,有25个临床症状(而不仅仅是4个)和2713行源数据。
我有一个Python代码使用熊猫库工作,但太慢,因为我使用嵌套的for-循环,和原始数据变得更重要。
# initial dataframe: return 9 rows
df = pd.DataFrame([
['PAT001','2022-01-23',1,2,0,0,2,3,3,1],
['PAT001','2022-01-24',1,2,0,0,2,3,3,1],
['PAT001','2022-01-25',1,2,0,0,2,3,3,1],
['PAT002','2022-01-23',1,2,0,0,2,3,3,1],
['PAT002','2022-01-24',1,2,0,0,2,3,3,1],
['PAT002','2022-01-25',1,2,0,0,2,3,3,1],
['PAT002','2022-01-26',1,2,0,0,2,3,3,1],
], columns=['pat_ide','sym_dat','sym_tou','sym_gra','sym_ete','sym_fie','sym_tou_del','sym_gra_del','sym_ete_del','sym_fie_del'])
tmp_df_exams = pd.DataFrame(columns = ['pat_ide','DOMAIN','USUBJID','FAOBJ','SYM_DAT','DEL'])
# list of 4 monitored clinical signs
signs = {
'sym_tou' : 'Dry cough',
'sym_gra' : 'Fatty cough',
'sym_ete' : 'Sneezing',
'sym_fie' : 'Fever'
}
for index, row in df.iterrows():
for k,v in signs.items():
tmp_df_exams = tmp_df_exams.append({
'pat_ide': row['pat_ide'],
'DOMAIN' : 'FA',
'USUBJID' : Patient_icon_format(row['pat_ide']),
'FAOBJ' : v,
'SYM_DAT' : row['sym_dat'],
'DEL' : row[f'{k}_del']
},ignore_index=True)目前,整个原始数据的结束需要几个小时。要想有一个想法,如果我筛选初始数据(9行),它需要在68s到81 s之间。
我怎样才能提高性能?
输出
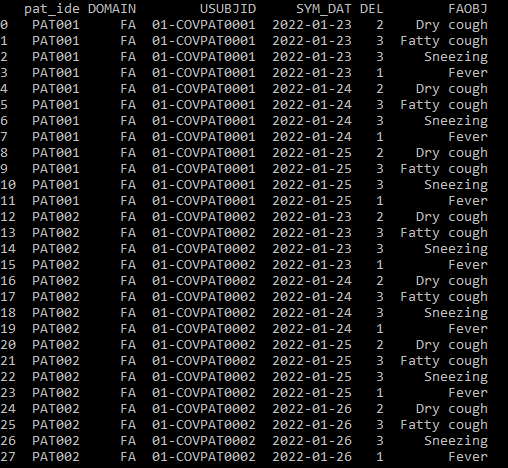
回答 1
Stack Overflow用户
发布于 2022-01-26 17:02:20
策略:构建多级列索引以更容易地操作数据
- 为列名创建元组映射
columns_map = dict()表示e在排序(df.columns):e_header = e.replace('_del',‘)如果e_header在符号:元组= (signse_header,'del’if‘del’if‘del’) columns_mape = tuple >>> columns_map {'sym_ete':(‘喷嚏’,'sym'),‘sym’,‘sym_ete_del:(’喷嚏‘,'del'),“sym_fie”:(“发烧”,“sym”),“sym_fie_del”:(“发烧”,“del”),“sym_gra”:(“脂肪性咳嗽”,“sym”),“sym_gra_del”:(“脂肪咳嗽”,“del”),“sym_tou”:(“干咳”,“sym”),“sym_tou_del”:(‘干咳’,'del')}
- 构建多级数据框架
dfh = df.set_index('pat_ide',‘sym_dat’).rename(列= columns_map) dfh.columns = pd.MultiIndex.from_tuples(dfh.columns,name=‘FAOBJ’),( data_name ) dfh = dfh.sort_index(axis=1) >>>dfh FAOBJ干咳脂肪性咳嗽发烧打喷嚏data_name del sym pat_ide sym_dat PAT001 2022-01-23 2 1 3 2 2 10.
- 数据杂耍
dfh = (dfh.swaplevel(axis=1) # put data_name level a)顶级.drop('sym',axis=1) # drop (未使用的数据).droplevel(‘data_name,axis=1)# level 0现在仅为’DEL‘.stack() #行到列.rename('DEL’),axis=1) #重命名数据.reset_index() #摆脱了多级索引)
- 插入缺失数据和类型
dfh = dfh.rename(columns={"sym_dat":"SYM_DAT"}) dfh'DOMAIN‘= 'FA’dfh'USUBJID‘= '01-COV'+ dfh'pat_ide’#模拟dfh = dfh.reindex('pat_ide',‘sym_dat’域‘,'USUBJID','FAOBJ','SYM_DAT','DEL',axis=1)
结果:
>>> dfh
pat_ide DOMAIN USUBJID FAOBJ SYM_DAT DEL
0 PAT001 FA 01-COVPAT001 Dry cough 2022-01-23 2
1 PAT001 FA 01-COVPAT001 Fatty cough 2022-01-23 3
2 PAT001 FA 01-COVPAT001 Fever 2022-01-23 1
3 PAT001 FA 01-COVPAT001 Sneezing 2022-01-23 3
4 PAT001 FA 01-COVPAT001 Dry cough 2022-01-24 2
5 PAT001 FA 01-COVPAT001 Fatty cough 2022-01-24 3
6 PAT001 FA 01-COVPAT001 Fever 2022-01-24 1
7 PAT001 FA 01-COVPAT001 Sneezing 2022-01-24 3
8 PAT001 FA 01-COVPAT001 Dry cough 2022-01-25 2
9 PAT001 FA 01-COVPAT001 Fatty cough 2022-01-25 3
10 PAT001 FA 01-COVPAT001 Fever 2022-01-25 1
11 PAT001 FA 01-COVPAT001 Sneezing 2022-01-25 3
12 PAT002 FA 01-COVPAT002 Dry cough 2022-01-23 2
13 PAT002 FA 01-COVPAT002 Fatty cough 2022-01-23 3
14 PAT002 FA 01-COVPAT002 Fever 2022-01-23 1
15 PAT002 FA 01-COVPAT002 Sneezing 2022-01-23 3
16 PAT002 FA 01-COVPAT002 Dry cough 2022-01-24 2
17 PAT002 FA 01-COVPAT002 Fatty cough 2022-01-24 3
18 PAT002 FA 01-COVPAT002 Fever 2022-01-24 1
19 PAT002 FA 01-COVPAT002 Sneezing 2022-01-24 3
20 PAT002 FA 01-COVPAT002 Dry cough 2022-01-25 2
21 PAT002 FA 01-COVPAT002 Fatty cough 2022-01-25 3
22 PAT002 FA 01-COVPAT002 Fever 2022-01-25 1
23 PAT002 FA 01-COVPAT002 Sneezing 2022-01-25 3
24 PAT002 FA 01-COVPAT002 Dry cough 2022-01-26 2
25 PAT002 FA 01-COVPAT002 Fatty cough 2022-01-26 3
26 PAT002 FA 01-COVPAT002 Fever 2022-01-26 1
27 PAT002 FA 01-COVPAT002 Sneezing 2022-01-26 3https://stackoverflow.com/questions/70849048
复制相似问题

