使用平滑与stat_summary均值和值相关的点大小。
使用平滑与stat_summary均值和值相关的点大小。
提问于 2022-01-09 10:44:29
我想就不同地点的土壤深度绘制土壤有机体的活动图。我将我的数据从宽到长的格式进行了重组,并创建了一个新的数据框架,其中只有三个必需变量(组1=土壤深度,组2=站点名称,x=土壤有机体活动的平均值)。我绘制的数据如下:
heatmap_plot <- ggplot(baits.activity.means, aes(x= Group.2, y=Group.1, size = x,
colour=Group.2))
heatmap_plot + stat_summary(fun = mean, geom = "point") +
labs(title = "Soil Biological Activity Distribution",
x = "Site", y = "Soil Depth") +
theme(plot.title = element_text(hjust = 0.5, face="bold")) +
scale_fill_discrete(name="Site")这使我有:
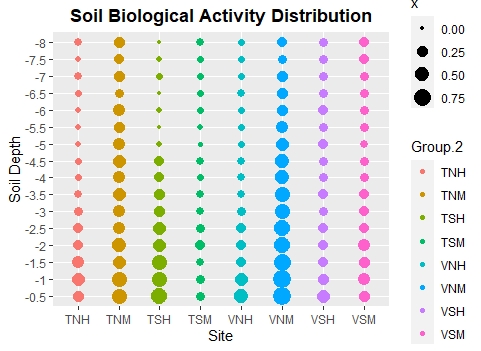
现在我为两件事而奋斗:
- ,首先,我希望y轴以递增的顺序显示土壤深度,所以从-8厘米到-0.5厘米(-8厘米是最低的土壤“层”)
- 编辑1:是否有一种简单的方法可以用相同的因子改变所有点的大小?
编辑2:我想出了另一种方法来说明我的数据,但我对如何开始感到迷茫。下面是一个粗略的草图:
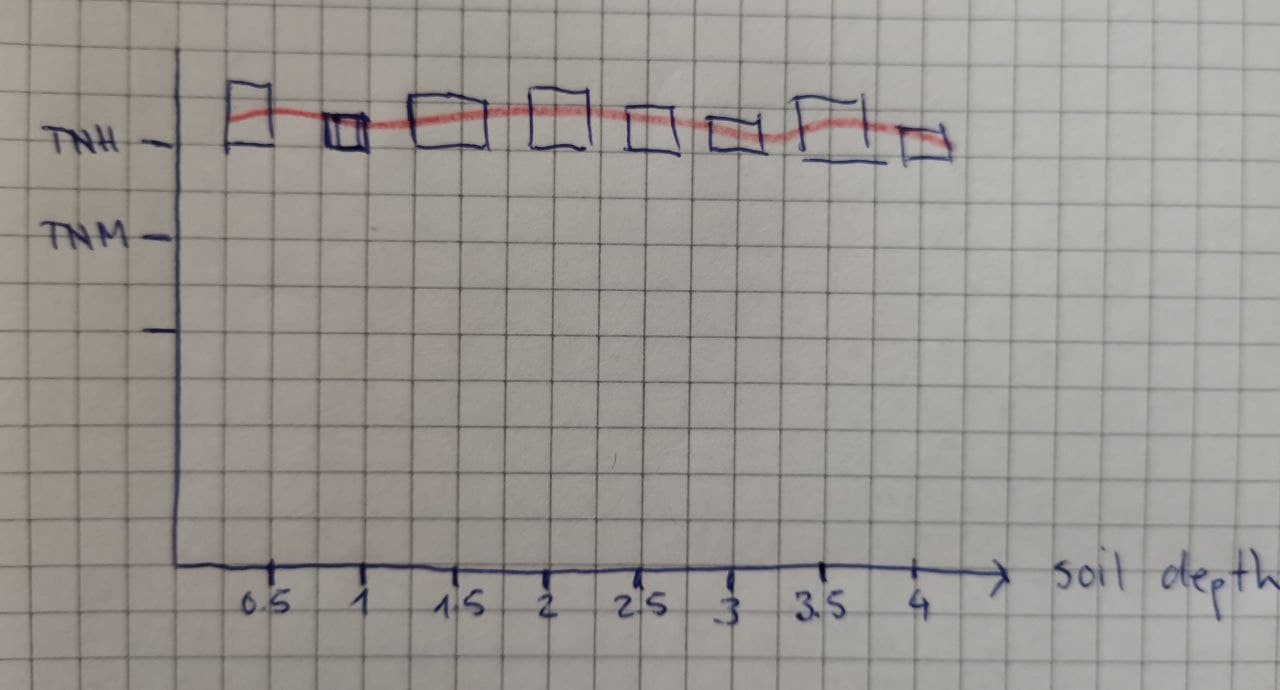
X轴表示土壤深度。对于每个土壤深度级别,我想用一个小节来表示平均活动值( bar越高,平均值越高)。理想情况下,我想添加一个平滑的显示沿土壤深度(x轴)的方法的过程。或者(但可能更复杂一些)来说明高值的条子,例如绿色的,红色的低值的条子--即使用颜色梯度来填充条子,以便更容易发现具有较高平均活度的土壤深度。
原始的长格式数据有超过5400个观测值,所以我只提供了使用聚合()收集的方法来提供较小的数据帧(baits.activity.means):
structure(list(Group.1 = structure(c(1L, 2L, 3L, 4L, 5L, 6L,
7L, 8L, 9L, 10L, 11L, 12L, 13L, 14L, 15L, 16L, 1L, 2L, 3L, 4L,
5L, 6L, 7L, 8L, 9L, 10L, 11L, 12L, 13L, 14L, 15L, 16L, 1L, 2L,
3L, 4L, 5L, 6L, 7L, 8L, 9L, 10L, 11L, 12L, 13L, 14L, 15L, 16L,
1L, 2L, 3L, 4L, 5L, 6L, 7L, 8L, 9L, 10L, 11L, 12L, 13L, 14L,
15L, 16L, 1L, 2L, 3L, 4L, 5L, 6L, 7L, 8L, 9L, 10L, 11L, 12L,
13L, 14L, 15L, 16L, 1L, 2L, 3L, 4L, 5L, 6L, 7L, 8L, 9L, 10L,
11L, 12L, 13L, 14L, 15L, 16L, 1L, 2L, 3L, 4L, 5L, 6L, 7L, 8L,
9L, 10L, 11L, 12L, 13L, 14L, 15L, 16L, 1L, 2L, 3L, 4L, 5L, 6L,
7L, 8L, 9L, 10L, 11L, 12L, 13L, 14L, 15L, 16L), .Label = c("-0.5",
"-1", "-1.5", "-2", "-2.5", "-3", "-3.5", "-4", "-4.5", "-5",
"-5.5", "-6", "-6.5", "-7", "-7.5", "-8"), class = "factor"),
Group.2 = c("TNH", "TNH", "TNH", "TNH", "TNH", "TNH", "TNH",
"TNH", "TNH", "TNH", "TNH", "TNH", "TNH", "TNH", "TNH", "TNH",
"TNM", "TNM", "TNM", "TNM", "TNM", "TNM", "TNM", "TNM", "TNM",
"TNM", "TNM", "TNM", "TNM", "TNM", "TNM", "TNM", "TSH", "TSH",
"TSH", "TSH", "TSH", "TSH", "TSH", "TSH", "TSH", "TSH", "TSH",
"TSH", "TSH", "TSH", "TSH", "TSH", "TSM", "TSM", "TSM", "TSM",
"TSM", "TSM", "TSM", "TSM", "TSM", "TSM", "TSM", "TSM", "TSM",
"TSM", "TSM", "TSM", "VNH", "VNH", "VNH", "VNH", "VNH", "VNH",
"VNH", "VNH", "VNH", "VNH", "VNH", "VNH", "VNH", "VNH", "VNH",
"VNH", "VNM", "VNM", "VNM", "VNM", "VNM", "VNM", "VNM", "VNM",
"VNM", "VNM", "VNM", "VNM", "VNM", "VNM", "VNM", "VNM", "VSH",
"VSH", "VSH", "VSH", "VSH", "VSH", "VSH", "VSH", "VSH", "VSH",
"VSH", "VSH", "VSH", "VSH", "VSH", "VSH", "VSM", "VSM", "VSM",
"VSM", "VSM", "VSM", "VSM", "VSM", "VSM", "VSM", "VSM", "VSM",
"VSM", "VSM", "VSM", "VSM"), x = c(0.230357142857143, 0.351785714285714,
0.282142857142857, 0.189285714285714, 0.169642857142857,
0.139285714285714, 0.0821428571428571, 0.0410714285714286,
0.0553571428571429, 0.0178571428571429, 0.0303571428571429,
0.0303571428571429, 0.0571428571428571, 0.0642857142857143,
0.0160714285714286, 0.0803571428571429, 0.61, 0.602222222222222,
0.414444444444444, 0.488888888888889, 0.337777777777778,
0.304444444444444, 0.366666666666667, 0.313333333333333,
0.348888888888889, 0.307777777777778, 0.254444444444444,
0.276666666666667, 0.124444444444444, 0.235555555555556,
0.165555555555556, 0.255555555555556, 0.664634146341463,
0.585365853658537, 0.535365853658537, 0.413414634146341,
0.395121951219512, 0.209756097560976, 0.252439024390244,
0.197560975609756, 0.170731707317073, 0.024390243902439,
0, 0.0146341463414634, 0, 0.0365853658536585, 0.0182926829268293,
0, 0.11, 0.143333333333333, 0.0944444444444444, 0.15, 0.127777777777778,
0.0666666666666667, 0.0744444444444444, 0.0555555555555556,
0.0755555555555556, 0.00555555555555556, 0.03, 0.05, 0.0511111111111111,
0.0611111111111111, 0.0555555555555556, 0.0888888888888889,
0.5, 0.346666666666667, 0.19, 0.191111111111111, 0.147777777777778,
0.1, 0.0988888888888889, 0.0866666666666667, 0.08, 0.0344444444444444,
0.0522222222222222, 0.0344444444444444, 0.0633333333333333,
0.0377777777777778, 0.0588888888888889, 0.0577777777777778,
0.911111111111111, 0.851111111111111, 0.75, 0.628888888888889,
0.644444444444444, 0.536666666666667, 0.541111111111111,
0.491111111111111, 0.425555555555556, 0.411111111111111,
0.29, 0.224444444444444, 0.232222222222222, 0.263333333333333,
0.13, 0.181111111111111, 0.377777777777778, 0.257777777777778,
0.198888888888889, 0.151111111111111, 0.147777777777778,
0.13, 0.168888888888889, 0.108888888888889, 0.106666666666667,
0.147777777777778, 0.126666666666667, 0.152222222222222,
0.125555555555556, 0.12, 0.101111111111111, 0.11, 0.262222222222222,
0.296666666666667, 0.241111111111111, 0.283333333333333,
0.198888888888889, 0.168888888888889, 0.15, 0.122222222222222,
0.11, 0.0877777777777778, 0.11, 0.111111111111111, 0.127777777777778,
0.127777777777778, 0.154444444444444, 0.15)), row.names = c(NA,
-128L), class = "data.frame")回答 1
Stack Overflow用户
发布于 2022-01-09 11:41:22
编辑:再想一想,小提琴的情节可能就是你想要的。它可以显示每个站点的值的分布。通过添加geom_violin(),您可以在ggplot中使用它。可能是个不错的开始。
这应该可以反转点1的y轴(见最后一行)。但我不能百分之百肯定你要的是第二点。
heatmap_plot <- ggplot(baits.activity.means, aes(x= Group.2, y=Group.1, size = x,
colour=Group.2))
heatmap_plot + stat_summary(fun = mean, geom = "point") +
labs(title = "Soil Biological Activity Distribution",
x = "Site", y = "Soil Depth") +
theme(plot.title = element_text(hjust = 0.5, face="bold")) +
scale_fill_discrete(name="Site") +
scale_y_discrete(limits = rev(levels(baits.activity.means$`Group.1`)))页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/70640496
复制相关文章
相似问题

