训练,调优,交叉验证和测试游侠(随机森林)分位数回归模型?
训练,调优,交叉验证和测试游侠(随机森林)分位数回归模型?
提问于 2022-01-04 18:43:00
有人可以分享如何训练,调优(超参数),交叉验证,并测试一个游侠分位数回归模型,以及误差评估?有虹膜或者波士顿的房产数据吗?
我之所以问这个问题,是因为我没有在Kaggle,随机博客,Youtube上找到很多使用分位数回归的例子或演练。我遇到的大多数问题都是分类问题。
我目前正在使用分位数回归模型,但我希望看到其他例子,特别是在超参数调优方面。
回答 1
Stack Overflow用户
回答已采纳
发布于 2022-01-05 01:32:31
这个函数有很多参数。因为这不是一个论坛,这一切意味着什么,我真的建议你点击交叉验证的问题,如何和为什么。(或者寻找可能已经回答的问题。)
library(tidyverse)
library(ranger)
library(caret)
library(funModeling)
data(iris)
#----------- setup data -----------
# this doesn't include exploration or cleaning which are both necessary
summary(iris)
df_status(iris)
#----------------- create training sample ----------------
set.seed(395280469) # for replicability
# create training sample partition (70/20 split)
tr <- createDataPartition(iris$Species,
p = .8,
list = F)有很多方法来分割数据,但我倾向于使用Caret,因为如果您需要这样做的话,它们就可以消除各种因素。
#--------- First model ---------
fit.r <- ranger(Sepal.Length ~ .,
data = iris[tr, ],
write.forest = TRUE,
importance = 'permutation',
quantreg = TRUE,
keep.inbag = TRUE,
replace = FALSE)
fit.r
# Ranger result
#
# Call:
# ranger(Sepal.Length ~ ., data = iris[tr, ], write.forest = TRUE,
# importance = "permutation", quantreg = TRUE, keep.inbag = TRUE,
# replace = FALSE)
#
# Type: Regression
# Number of trees: 500
# Sample size: 120
# Number of independent variables: 4
# Mtry: 2
# Target node size: 5
# Variable importance mode: permutation
# Splitrule: variance
# OOB prediction error (MSE): 0.1199364
# R squared (OOB): 0.8336928
p.r <- predict(fit.r, iris[-tr, -1],
type = 'quantiles')它默认为.1、.5和.9:
postResample(p.r$predictions[, 1], iris[-tr, 1])
# RMSE Rsquared MAE
# 0.5165946 0.7659124 0.4036667
postResample(p.r$predictions[, 2], iris[-tr, 1])
# RMSE Rsquared MAE
# 0.3750556 0.7587326 0.3133333
postResample(p.r$predictions[, 3], iris[-tr, 1])
# RMSE Rsquared MAE
# 0.6488991 0.7461830 0.5703333 要了解这在实践中的样子:
# this performance is the best so far, let's see what it looks like visually
ggplot(data.frame(p.Q1 = p.r$predictions[, 1],
p.Q5 = p.r$predictions[, 2],
p.Q9 = p.r$predictions[, 3],
Actual = iris[-tr, 1])) +
geom_point(aes(x = Actual, y = p.Q1, color = "P.Q1")) +
geom_point(aes(x = Actual, y = p.Q5, color = "P.Q5")) +
geom_point(aes(x = Actual, y = p.Q9, color = "P.Q9")) +
geom_line(aes(Actual, Actual, color = "Actual")) +
scale_color_viridis_d(end = .8, "Error",
direction = -1)+
theme_bw()
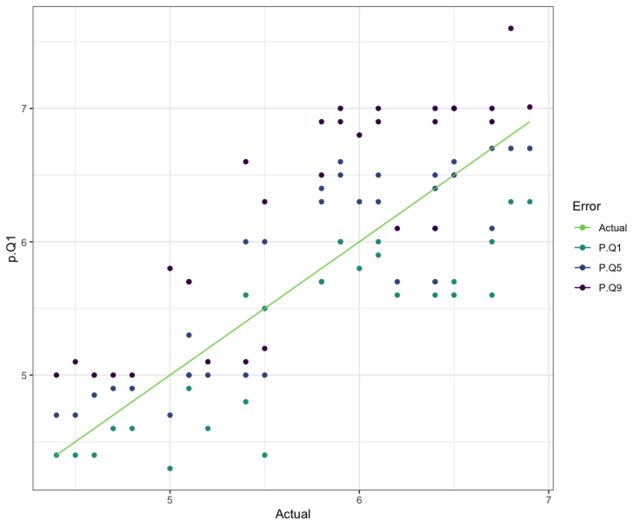
# since Quantile .1 performed the best
ggplot(data.frame(p.Q9 = p.r$predictions[, 3],
Actual = iris[-tr, 1])) +
geom_point(aes(x = Actual, y = p.Q9, color = "P.Q9")) +
geom_segment(aes(x = Actual, xend = Actual,
y = Actual, yend = p.Q9)) +
geom_line(aes(Actual, Actual, color = "Actual")) +
scale_color_viridis_d(end = .8, "Error",
direction = -1)+
theme_bw()
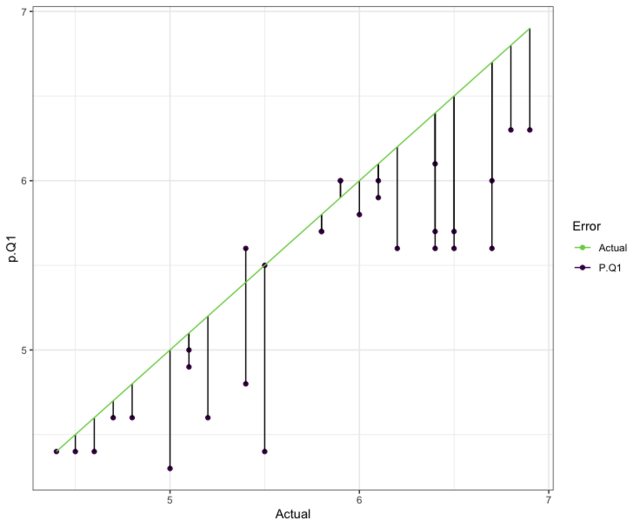
#------------ ranger model with options --------------
# last call used default
# splitrule: variance, use "extratrees" (only 2 for this one)
# mtry = 2, use 3 this time
# min.node.size = 5, using 6 this time
# using num.threads = 15 ** this is the number of cores on YOUR device
# change accordingly --- if you don't know, drop this one
set.seed(326)
fit.r2 <- ranger(Sepal.Length ~ .,
data = iris[tr, ],
write.forest = TRUE,
importance = 'permutation',
quantreg = TRUE,
keep.inbag = TRUE,
replace = FALSE,
splitrule = "extratrees",
mtry = 3,
min.node.size = 6,
num.threads = 15)
fit.r2
# Ranger result
# Type: Regression
# Number of trees: 500
# Sample size: 120
# Number of independent variables: 4
# Mtry: 3
# Target node size: 6
# Variable importance mode: permutation
# Splitrule: extratrees
# Number of random splits: 1
# OOB prediction error (MSE): 0.1107299
# R squared (OOB): 0.8464588 这种模式也产生了类似的结果。
p.r2 <- predict(fit.r2, iris[-tr, -1],
type = 'quantiles')
postResample(p.r2$predictions[, 1], iris[-tr, 1])
# RMSE Rsquared MAE
# 0.4932883 0.8144309 0.4000000
postResample(p.r2$predictions[, 2], iris[-tr, 1])
# RMSE Rsquared MAE
# 0.3610171 0.7643744 0.3100000
postResample(p.r2$predictions[, 3], iris[-tr, 1])
# RMSE Rsquared MAE
# 0.6555939 0.8141144 0.5603333 总体上来说,这一预测也相当相似。这不是一组非常大的数据,没有几个预测器。他们贡献了多少?
importance(fit.r2)
# Sepal.Width Petal.Length Petal.Width Species
# 0.06138883 0.71052453 0.22956522 0.18082998
#------------ ranger model with options --------------
# drop a predictor, lower mtry, min.node.size
set.seed(326)
fit.r3 <- ranger(Sepal.Length ~ .,
data = iris[tr, -4], # dropped Sepal.Width
write.forest = TRUE,
importance = 'permutation',
quantreg = TRUE,
keep.inbag = TRUE,
replace = FALSE,
splitrule = "extratrees",
mtry = 2, # has to change (var count lower)
min.node.size = 4, # lowered
num.threads = 15)
fit.r3
# Ranger result
# Type: Regression
# Number of trees: 500
# Sample size: 120
# Number of independent variables: 3
# Mtry: 2
# Target node size: 6
# Variable importance mode: permutation
# Splitrule: extratrees
# Number of random splits: 1
# OOB prediction error (MSE): 0.1050143
# R squared (OOB): 0.8543842 第二种最重要的预测因子被移除并得到改善。
p.r3 <- predict(fit.r3, iris[-tr, -c(1, 4)],
type = 'quantiles')
postResample(p.r3$predictions[, 1], iris[-tr, 1])
# RMSE Rsquared MAE
# 0.4760952 0.8089810 0.3800000
postResample(p.r3$predictions[, 2], iris[-tr, 1])
# RMSE Rsquared MAE
# 0.3738315 0.7769388 0.3250000
postResample(p.r3$predictions[, 3], iris[-tr, 1])
# RMSE Rsquared MAE
# 0.6085584 0.8032592 0.5170000
importance(fit.r3)
# almost everthing relies on Petal.Length
# Sepal.Width Petal.Length Species
# 0.08008264 0.95440333 0.32570147 页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/70583626
复制相关文章
相似问题

