如何用R中的序号来更改对象的名称?
如何用R中的序号来更改对象的名称?
提问于 2022-01-04 14:13:10
我有一个数据文件,其中包含了像P95、P104等编码的基因。这个数字反映了基因名称列表中的顺序。
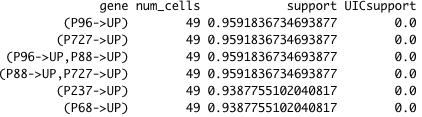
基因名称清单(其中有2000份):

在这种情况下,如何将dataframe中的P##转换为基因名称?
UPD:这里有一个示例dataframe和一个基因列表:
gene <- c("(P10->UP)", "(P2->UP, P9->UP)", "(P10->UP, P3->UP)", "(P5->NORM, P7->UP)")
support <- c(0.95, 0.94, 0.93, 0.92)
df <- data.frame(gene, support)
gene_list <- c("A", "B", "C", "D", "E", "F", "G", "H", "I", "J")P10与第10基因"J“相对应,P2为"B”等。
我想得到的结果应该如下所示:

回答 1
Stack Overflow用户
回答已采纳
发布于 2022-01-04 21:06:15
一种方法可能是使用来自mutate包的separate和tidyverse包。
separate不能将列拆分为未知的列数。因此,我不得不计算基因列优先(max_genes)中的最大基因数。
数据
gene <- c("(P10->UP)", "(P2->UP, P9->UP)", "(P10->UP, P3->UP)", "(P5->NORM, P7->UP)")
support <- c(0.95, 0.94, 0.93, 0.92)
df <- data.frame(gene, support)
gene_list <- c("A", "B", "C", "D", "E", "F", "G", "H", "I", "J")码
# calculate max number of genes in column gene for spreading
max_genes = ncol(str_extract_all(df$gene, "->", simplify = T))
df %>%
# remove brackets and spaces in column gene
mutate(gene = str_remove_all(gene, "[(|)|\\s]")) %>%
# separate gene into name and expresssion
separate(col = gene,
sep = "->|,",
into = paste0(c("gene_name", "exp"),
rep(1:max_genes, each = 2)),
fill = "right") %>%
# substitute gene number with gene name
mutate(across(starts_with("gene_name"), ~gene_list[as.numeric(str_remove(., "P"))]))输出
gene_name1 exp1 gene_name2 exp2 support
1 J UP <NA> <NA> 0.95
2 B UP I UP 0.94
3 J UP C UP 0.93
4 E NORM G UP 0.92页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/70580102
复制相关文章
相似问题

