Plotnine:按列排列的面
Plotnine:按列排列的面
提问于 2021-12-19 09:59:58
我的dataframe MedComp有以下结构:
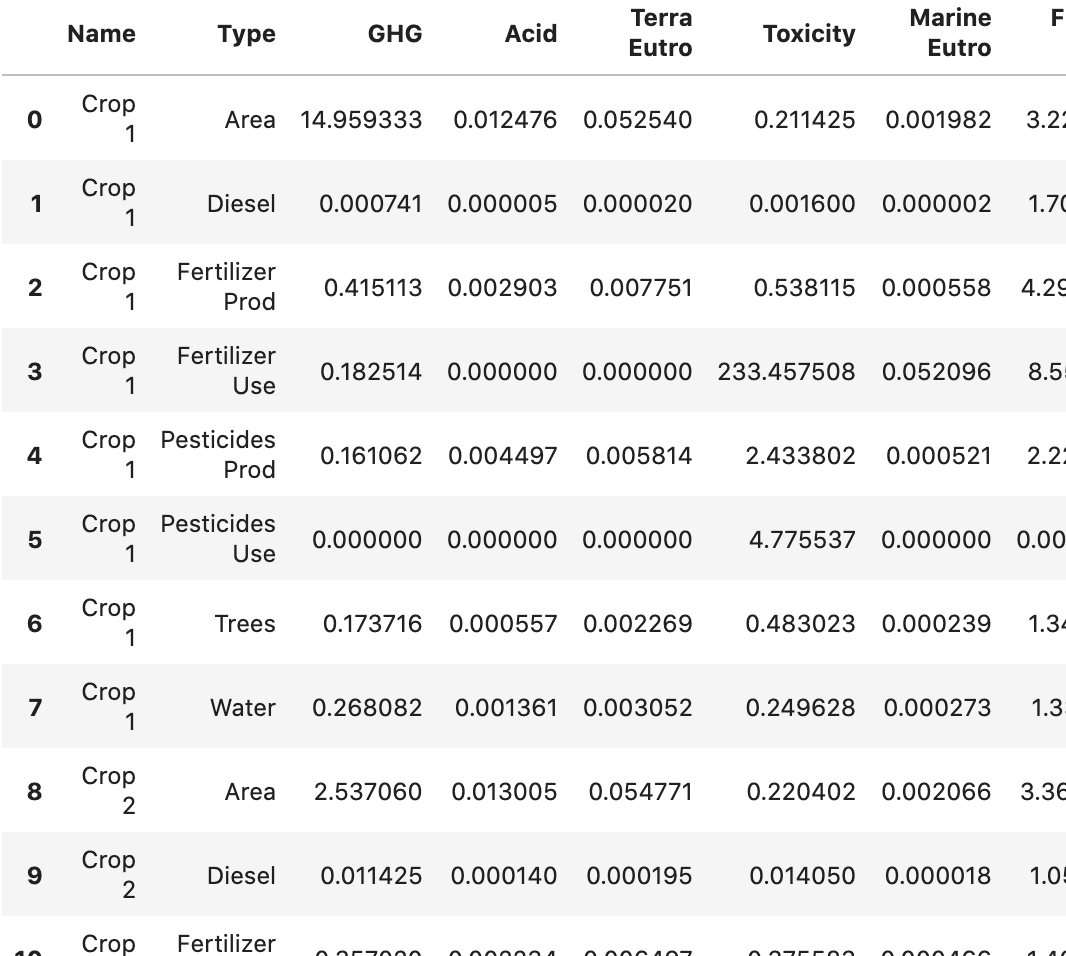
我现在想要做一个堆叠的条形图,每个Name一个(有两个不同的),由Type堆叠(目前使用颜色)。Y轴是其余的列,由于范围不同,每列都有一个单独的图。对于Name中的一种作物,我还想为总数显示一个错误栏,目前在一个名为FarmQuant的单独数据中计算,如下所示:
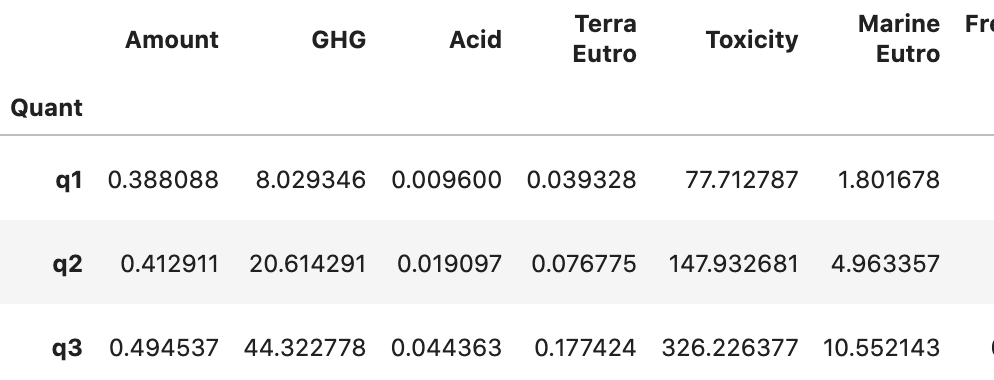
到目前为止,我只使用了一个循环,从而创建了每个列的一个数字:
Midpoint = [['GHG', 'Total climate change [kg CO2e]'],
['Acid', 'Freshwater & ter. acidification [mol H+-Eq]'],
['Terra Eutro', 'Terrestrial eutrophication [mol N-Eq]'],
['Toxicity', 'Freshwater ecotoxicity [CTU]'],
['Marine Eutro', 'Marine eutrophication [kg N-Eq]' ],
]
for MPID in range(0, len(Midpoint)):
print(MPID+1, len(Midpoint), ' ', end='')
q = (ggplot(Farm_Quant))
+ geom_col(MedComb, aes('Name', Midpoint[MPID][0], fill='Type'))
+ scale_fill_brewer(type='div', palette=2)
+ geom_point(Farm_Quant, aes(x=1, y=Farm_Quant.loc['q2',Midpoint[MPID][0]]))
+ geom_errorbar(aes(x=1,ymin = Farm_Quant.loc['q1',Midpoint[MPID][0]], ymax = Farm_Quant.loc['q3',Midpoint[MPID][0]]))
+ theme_matplotlib()
+ theme(figure_size=(2.2, 4), legend_position = (1.25, 0.5),
axis_title_x =element_blank(),
axis_ticks_major_x=element_blank())
+ scale_y_continuous(name=Midpoint[MPID][1])
+ labs(title = Midpoint[MPID][0])
)
fig = q.draw()
fig.show()然而,我想要这样做,使用方面,使所有在一个数字,只有一个传奇。有谁知道怎么用这个吗?
请只提供matplotlib (据我所知不支持面设置)或plot9的示例,因为我已经完成了其他几个情节,并且希望保持相同的外观。另外,这些也是我最熟悉的。
编辑:这里还有一些要处理的测试数据:
MedComb = pd.DataFrame({
'Name' : ['Crop1', 'Crop1', 'Crop1', 'Crop1', 'Crop2', 'Crop2', 'Crop2', 'Crop2'],
'Type' : ['Area', 'Diesel', 'Fert', 'Pest', 'Area', 'Diesel', 'Fert', 'Pest'],
'GHG': [14.9, 0.0007, 0.145, 0.1611, 2.537, 0.011, 0.1825, 0.115],
'Acid': [0.0125, 0.0005, 0.0029, 0.0044, 0.013, 0.00014, 0.0033, 0.0055],
'Terra Eutro': [0.053, 0.0002, 0.0077, 0.0001, 0.0547, 0.00019, 0.0058, 0.0002]
})
Farm_Quant = pd.DataFrame({
'Amount': [0.388, 0.4129, 0.1945],
'GHG': [8.029, 20.61, 44.32],
'Acid': [0.009, 0.019, 0.044],
'Terra Eutro': [0.039, 0.077, 0.0177]},
index = ['q1', 'q2', 'q3']
)回答 1
Stack Overflow用户
回答已采纳
发布于 2021-12-28 01:43:53
当您的geom美学在他们自己的专栏中时,Plotnine的效果最好,因此您可以通过使用melt来实现这一点。另外,如果您将Farm_Quant数据remove加入到您融化的MedComb数据文件中,那么您只需引用该单个数据并删除for循环即可。
# Sample Data
MedComb = pd.DataFrame({
'Name' : ['Crop1', 'Crop1', 'Crop1', 'Crop1', 'Crop2', 'Crop2', 'Crop2', 'Crop2'],
'Type' : ['Area', 'Diesel', 'Fert', 'Pest', 'Area', 'Diesel', 'Fert', 'Pest'],
'GHG': [14.9, 0.0007, 0.145, 0.1611, 2.537, 0.011, 0.1825, 0.115],
'Acid': [0.0125, 0.0005, 0.0029, 0.0044, 0.013, 0.00014, 0.0033, 0.0055],
'Terra Eutro': [0.053, 0.0002, 0.0077, 0.0001, 0.0547, 0.00019, 0.0058, 0.0002]
})
Farm_Quant = pd.DataFrame({
'Amount': [0.388, 0.4129, 0.1945],
'GHG': [8.029, 20.61, 44.32],
'Acid': [0.009, 0.019, 0.044],
'Terra Eutro': [0.039, 0.077, 0.0177]},
index = ['q1', 'q2', 'q3']
)
# melt the MedComb df from wide to long
med_comb_long = pd.melt(MedComb,id_vars=['Name','Type'],
var_name='midpoint',value_name='value')
# Basically transpose the Farm_Quant df in two steps to join with med_comb_long
# convert the Farm_Quant df from wide to long and make the quarters a column
midpoint_long = pd.melt(Farm_Quant.reset_index().drop(columns=['Amount']),
id_vars=['index'],var_name='midpoint',value_name='error_bar')
# make long farm quant df wide again, but with the quarters as the columns and facet names as index
midpoint_reshape = pd.pivot(midpoint_long,index='midpoint',columns='index',
values='error_bar')
# Join the data for the bar charts and error bars/points into single df
plot_data = med_comb_long.join(midpoint_reshape,on='midpoint')
# make the plot
q2 = (ggplot(plot_data)
+ geom_col(aes('Name','value',fill='Type'))
+ scale_fill_brewer(type='div', palette=2)
+ facet_wrap('midpoint',scales='free')
+ geom_errorbar(aes(x=1,ymin='q1',ymax='q3'))
+ geom_point(aes(x=1,y='q2'))
+ theme_matplotlib()
# + theme(figure_size=(2.2, 4), legend_position = (1.25, 0.5),
# axis_title_x =element_blank(),
# axis_ticks_major_x=element_blank())
+ scale_y_continuous(name=Midpoint[MPID][1])
+ labs(title = Midpoint[MPID][0])
)
q2
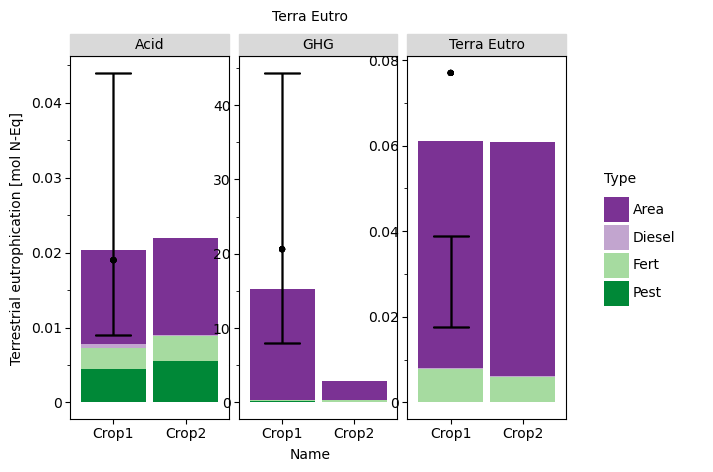
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/70410415
复制相关文章
相似问题

