微调蛋白质语言模型——protein level
微调蛋白质语言模型——protein level

Tom2Code
发布于 2026-04-17 17:29:36
发布于 2026-04-17 17:29:36
一.前言
今天给大家带来微调蛋白质语言模型的教程。
今天这个任务是针对
1.per protein prediction,这个就是说使用蛋白质语言模型对整个蛋白质序列进行全局层面的预测输出。
除了今天这一篇以外还会有两篇内容是关于:
2.per residue prediction
3.per residue regerssion
大家可以自己找自己任务匹配的数据集,然后微调一个属于自己的蛋白质语言模型。
二.背景
蛋白质大语言模型的微调的两个方法:
1.PEFT:Parameter Efficient Fine Tuning
https://huggingface.co/blog/peft
2.LoRA:Low-Rank Adaptation of large language models
https://arxiv.org/abs/2106.09685
我们今天使用 LoRA
所以简单的介绍一下LoRA的工作原理:
LoRA 不需要重新训练整个模型,而是按原样冻结模型的原始权重和参数。然后,在这个原始模型的基础上,它添加了一个名为低秩矩阵的轻量级附加组件,然后将其应用于新的输入,以获得特定于上下文的结果。低秩矩阵会对原始模型的权重进行调整,使输出结果与所需的用例相匹配。
LoRA 充分利用低秩矩阵的概念,使模型训练过程极其高效和快速。传统上,对 LLM 进行微调需要调整整个模型。LoRA 专注于修改小部分参数(低秩矩阵),以减少计算和内存开销。
该图显示了 LoRA 如何通过使用秩为 r 的较小矩阵来更新矩阵 A 和 B,以跟踪预训练权重的变化。LoRA 训练完成后,较小的权重会被合并到一个新的权重矩阵中,而无需修改预训练模型的原始权重。
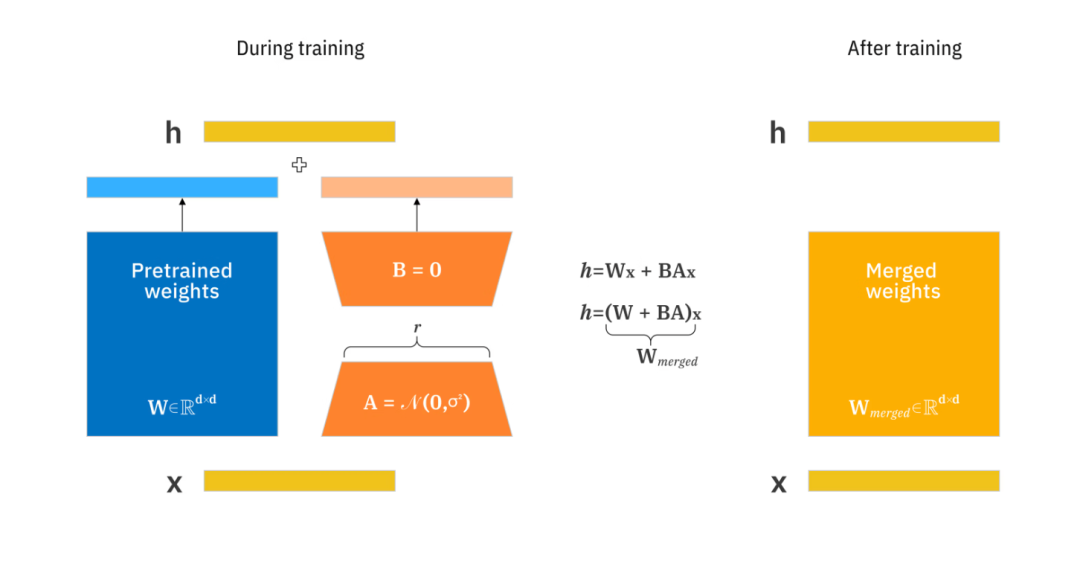
可以看到左侧的pretrained weights就是冻结的预训练的权重,这一部分将会被冻结,我们需要训练和学习的是右侧的两个倒装的梯形的权重。然后在最终使用的时候 我们将会把蓝色长方形的预训练的权重和梯形的权重融合在一起,进行预测。
LoRA 建立在这样的理解之上:大型模型本质上具有低维结构。通过利用称为低秩矩阵的较小矩阵,LoRA 可以有效地适应这些模型。该方法侧重于核心概念,即重大的模型变化可以用更少的参数来表示,从而提高适应过程的效率。
矩阵是机器学习和神经网络工作方式的重要组成部分。低秩矩阵比较大或较高秩矩阵更小,数值也更少。它们不会占用太多内存,并且需要更少的步骤来相加或相乘,使计算机处理速度更快。
一个高秩矩阵可以分解成两个低秩矩阵,一个 4 x 4 矩阵可以分解成一个 4 x 1 和一个 1 x 4 矩阵。
所以这里的低秩矩阵,也就是我们新的神经网络中需要学习的参数。
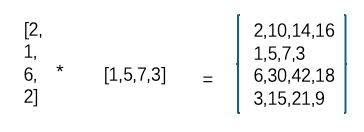
LoRA 将低秩矩阵添加到冻结的原始机器学习模型中。在微调期间,低秩矩阵通过梯度下降进行更新,而不修改基础模型的权重。这些矩阵包含在生成结果时应用于模型的新权重。乘法变化矩阵与基础模型权重相加,得到最终的微调模型。该过程以最小的计算能力和训练时间改变模型产生的输出。
就本质而言,LoRA 保持原始模型不变,为模型的每一层添加可改变的小型部分。这大大减少了模型的可训练参数和训练过程所需的 GPU 内存,这对于微调或训练大型模型而言是另一个重大挑战。
so更详细的介绍大家可以去看LoRA的论文,接下来我们直接进入代码。
三.代码实战
首先说一下运行环境,这份代码运行在Google colab的T4 gpu上
其次本次我们是为了演示起来比较方便,我们使用esm2家族最小的蛋白质语言模型:esm-8m
关于esm 蛋白质语言模型家族的介绍:
https://github.com/facebookresearch/esm
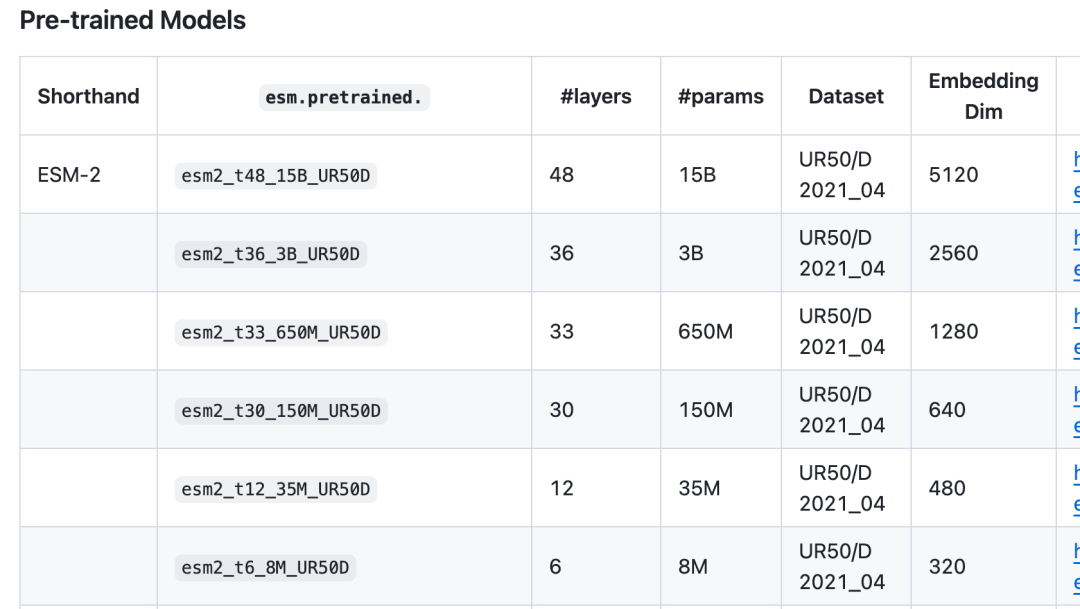
读者朋友们还可以微调其他模型:

因为一开始就说了这是一个per protein prediction的任务,我们本次的任务是直接对整个蛋白质的sequence做一个预测,所以数据集也是对应的类型。
我们用到的是gb1数据集,这是一个蛋白质序列和fitness对应的数据集,包含16万种GB1变体的经典“序列-功能(fitness)”回归数据集,完美保留了真实的突变上位效应,极其适合用来评估AI模型能否从简单的低阶突变中精准外推预测高阶复杂突变的适应度。
数据集地址:
https://github.com/J-SNACKKB/FLIP/tree/main/splits/gb1
好了接下来直接开始代码,因为时间有限,大家代码有任何问题欢迎讨论交流,没有写很详细的注释,如果需要可以借助大语言模型进行加深理解:
3.1依赖安装:
!pip install datasets
!pip install evaluate
!pip install SentencePiece
!pip install transformers[torch]
!pip install peft
!pip3 install deepspeed
3.2依赖导入
import os.path
from google.colab import drive
drive.mount('/content/drive')
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.nn import BCEWithLogitsLoss, CrossEntropyLoss, MSELoss
from torch.utils.data import DataLoader
import re
import numpy as np
import pandas as pd
import copy
import transformers, datasets
from transformers.modeling_outputs import SequenceClassifierOutput
from transformers.models.t5.modeling_t5 import T5Config, T5PreTrainedModel, T5Stack
#from transformers.utils.model_parallel_utils import assert_device_map, get_device_map
from transformers import T5EncoderModel, T5Tokenizer
from transformers import EsmModel, AutoTokenizer, AutoModelForSequenceClassification
from transformers import TrainingArguments, Trainer, set_seed
import peft
from peft import get_peft_config, PeftModel, PeftConfig, inject_adapter_in_model, LoraConfig
from evaluate import load
from datasets import Dataset
from tqdm import tqdm
import random
from scipy import stats
from sklearn.metrics import accuracy_score
import matplotlib.pyplot as plt3.3环境版本,配置deepspeed
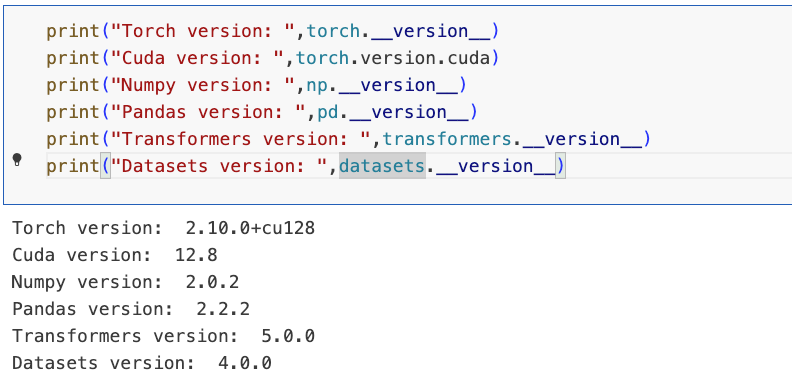
# Set environment variables to run Deepspeed from a notebook
os.environ["MASTER_ADDR"] = "localhost"
os.environ["MASTER_PORT"] = "9994" # modify if RuntimeError: Address already in use
os.environ["RANK"] = "0"
os.environ["LOCAL_RANK"] = "0"
os.environ["WORLD_SIZE"] = "1"
3.4模型列表
ESMs = ["facebook/esm2_t6_8M_UR50D",
"facebook/esm2_t12_35M_UR50D",
"facebook/esm2_t30_150M_UR50D",
"facebook/esm2_t33_650M_UR50D",
"facebook/esm2_t36_3B_UR50D"]
T5s = ["Rostlab/prot_t5_xl_uniref50",
'Rostlab/ProstT5',
"ElnaggarLab/ankh-base",
"ElnaggarLab/ankh-large"]
我们使用这个
checkpoint=ESMs[0]
3.5准备输入数据&划分数据集
# For this example we import the "three_vs_rest" GB1 dataset from https://github.com/J-SNACKKB/FLIP
# For details, see publication here: https://openreview.net/forum?id=p2dMLEwL8tF
import requests
import zipfile
from io import BytesIO
# Download the zip file from GitHub
url = 'https://github.com/J-SNACKKB/FLIP/raw/main/splits/gb1/splits.zip'
response = requests.get(url)
zip_file = zipfile.ZipFile(BytesIO(response.content))
# Load the `three_vs_rest.csv` file into a pandas dataframe
with zip_file.open('splits/three_vs_rest.csv') as file:
df = pd.read_csv(file)
# Drop test data
df=df[df.set=="train"]
# Get train and validation data
my_train=df[df.validation!=True].reset_index(drop=True)
my_valid=df[df.validation==True].reset_index(drop=True)
# Set column names to "sequence" and "label"
my_train.columns=["sequence","label"]+list(my_train.columns[2:])
my_valid.columns=["sequence","label"]+list(my_valid.columns[2:])
# Drop unneeded columns
my_train=my_train[["sequence","label"]]
my_valid=my_valid[["sequence","label"]]
my_train.head(5)
输出:
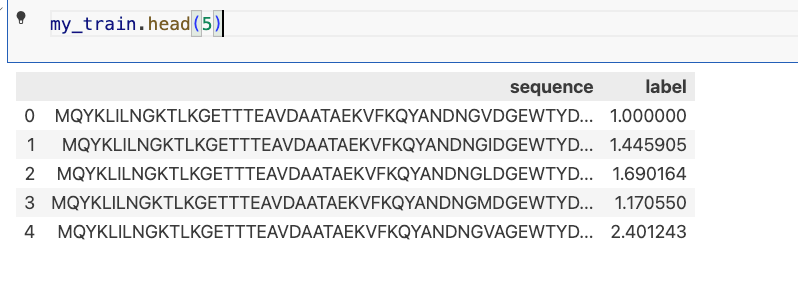
3.6准备esm加载类
#load ESM2 models
def load_esm_model(checkpoint, num_labels, half_precision, full = False, deepspeed=True):
tokenizer = AutoTokenizer.from_pretrained(checkpoint)
if half_precision and deepspeed:
model = AutoModelForSequenceClassification.from_pretrained(checkpoint, num_labels=num_labels, torch_dtype=torch.float16)
else:
model = AutoModelForSequenceClassification.from_pretrained(checkpoint, num_labels=num_labels)
if full == True:
return model, tokenizer
peft_config = LoraConfig(
r=4, lora_alpha=1, bias="all", target_modules=["query","key","value","dense"]
)
model = inject_adapter_in_model(peft_config, model)
# Unfreeze the prediction head
for (param_name, param) in model.classifier.named_parameters():
param.requires_grad = True
return model, tokenizer
3.7定义训练参数
# Deepspeed config for optimizer CPU offload
ds_config = {
"fp16": {
"enabled": "auto",
"loss_scale": 0,
"loss_scale_window": 1000,
"initial_scale_power": 16,
"hysteresis": 2,
"min_loss_scale": 1
},
"optimizer": {
"type": "AdamW",
"params": {
"lr": "auto",
"betas": "auto",
"eps": "auto",
"weight_decay": "auto"
}
},
"scheduler": {
"type": "WarmupLR",
"params": {
"warmup_min_lr": "auto",
"warmup_max_lr": "auto",
"warmup_num_steps": "auto"
}
},
"zero_optimization": {
"stage": 2,
"offload_optimizer": {
"device": "cpu",
"pin_memory": True
},
"allgather_partitions": True,
"allgather_bucket_size": 2e8,
"overlap_comm": True,
"reduce_scatter": True,
"reduce_bucket_size": 2e8,
"contiguous_gradients": True
},
"gradient_accumulation_steps": "auto",
"gradient_clipping": "auto",
"steps_per_print": 2000,
"train_batch_size": "auto",
"train_micro_batch_size_per_gpu": "auto",
"wall_clock_breakdown": False
}
# Set random seeds for reproducibility of your trainings run
def set_seeds(s):
torch.manual_seed(s)
np.random.seed(s)
random.seed(s)
set_seed(s)
# Dataset creation
def create_dataset(tokenizer,seqs,labels):
tokenized = tokenizer(seqs, max_length=1024, padding=True, truncation=True)
dataset = Dataset.from_dict(tokenized)
dataset = dataset.add_column("labels", labels)
return dataset
# Main training fuction
def train_per_protein(
checkpoint, #model checkpoint
train_df, #training data
valid_df, #validation data
num_labels = 1, #1 for regression, >1 for classification
# effective training batch size is batch * accum
# we recommend an effective batch size of 8
batch = 4, #for training
accum = 2, #gradient accumulation
val_batch = 16, #batch size for evaluation
epochs = 10, #training epochs
lr = 3e-4, #recommended learning rate
seed = 42, #random seed
deepspeed = False,#if gpu is large enough disable deepspeed for training speedup
mixed = True, #enable mixed precision training
full = False, #enable training of the full model (instead of LoRA)
gpu = 1 ): #gpu selection (1 for first gpu)
print("Model used:", checkpoint, "\n")
# Correct incompatible training settings
if"ankh"in checkpoint and mixed:
print("Ankh models do not support mixed precision training!")
print("switched to FULL PRECISION TRAINING instead")
mixed = False
# Set gpu device
os.environ["CUDA_VISIBLE_DEVICES"]=str(gpu-1)
# Set all random seeds
set_seeds(seed)
# load model
if"esm"in checkpoint:
model, tokenizer = load_esm_model(checkpoint, num_labels, mixed, full, deepspeed)
else:
model, tokenizer = load_T5_model(checkpoint, num_labels, mixed, full, deepspeed)
# Preprocess inputs
# Replace uncommon AAs with "X"
train_df["sequence"]=train_df["sequence"].str.replace('|'.join(["O","B","U","Z","J"]),"X",regex=True)
valid_df["sequence"]=valid_df["sequence"].str.replace('|'.join(["O","B","U","Z","J"]),"X",regex=True)
# Add spaces between each amino acid for ProtT5 and ProstT5 to correctly use them
if"Rostlab"in checkpoint:
train_df['sequence']=train_df.apply(lambda row : " ".join(row["sequence"]), axis = 1)
valid_df['sequence']=valid_df.apply(lambda row : " ".join(row["sequence"]), axis = 1)
# Add <AA2fold> for ProstT5 to inform the model of the input type (amino acid sequence here)
if"ProstT5"in checkpoint:
train_df['sequence']=train_df.apply(lambda row : "<AA2fold> " + row["sequence"], axis = 1)
valid_df['sequence']=valid_df.apply(lambda row : "<AA2fold> " + row["sequence"], axis = 1)
# Create Datasets
train_set=create_dataset(tokenizer,list(train_df['sequence']),list(train_df['label']))
valid_set=create_dataset(tokenizer,list(valid_df['sequence']),list(valid_df['label']))
# Huggingface Trainer arguments
args = TrainingArguments(
"./",
eval_strategy = "epoch",
logging_strategy = "epoch",
save_strategy = "no",
learning_rate=lr,
per_device_train_batch_size=batch,
per_device_eval_batch_size=val_batch,
gradient_accumulation_steps=accum,
num_train_epochs=epochs,
seed = seed,
deepspeed= ds_config if deepspeed else None,
fp16 = mixed,
)
# Metric definition for validation data
def compute_metrics(eval_pred):
if num_labels>1: # for classification
metric = load("accuracy")
predictions, labels = eval_pred
predictions = np.argmax(predictions, axis=1)
else: # for regression
metric = load("spearmanr")
predictions, labels = eval_pred
return metric.compute(predictions=predictions, references=labels)
# Trainer
trainer = Trainer(
model,
args,
train_dataset=train_set,
eval_dataset=valid_set,
processing_class=tokenizer,
compute_metrics=compute_metrics
)
# Train model
trainer.train()
return tokenizer, model, trainer.state.log_history
3.8开始微调模型
tokenizer, model, history = train_per_protein(checkpoint, my_train, my_valid, num_labels = 1, batch = 8, accum = 1, epochs = 50, seed = 42, mixed = True)
输出:
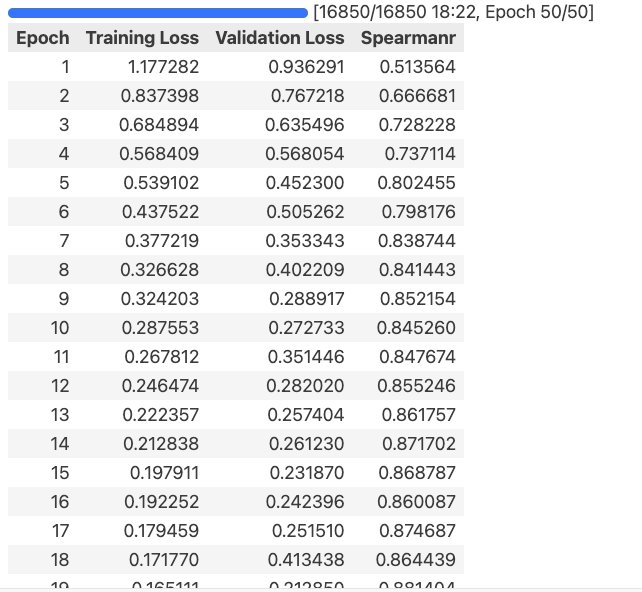
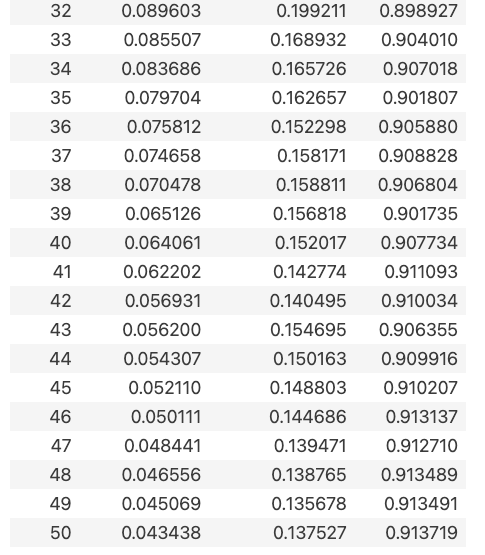
3.9可视化
# Get loss, val_loss, and the computed metric from history
loss = [x['loss'] for x inhistoryif'loss'in x]
val_loss = [x['eval_loss'] for x inhistoryif'eval_loss'in x]
# Get spearman (for regression) or accuracy value (for classification)
if [x['eval_spearmanr'] for x inhistoryif'eval_spearmanr'in x] != []:
metric = [x['eval_spearmanr'] for x inhistoryif'eval_spearmanr'in x]
else:
metric = [x['eval_accuracy'] for x inhistoryif'eval_accuracy'in x]
epochs = [x['epoch'] for x inhistoryif'loss'in x]
# Create a figure with two y-axes
fig, ax1 = plt.subplots(figsize=(10, 5))
ax2 = ax1.twinx()
# Plot loss and val_loss on the first y-axis
line1 = ax1.plot(epochs, loss, label='train_loss')
line2 = ax1.plot(epochs, val_loss, label='val_loss')
ax1.set_xlabel('Epoch')
ax1.set_ylabel('Loss')
# Plot the computed metric on the second y-axis
line3 = ax2.plot(epochs, metric, color='red', label='val_metric')
ax2.set_ylabel('Metric')
ax2.set_ylim([0, 1])
# Combine the lines from both y-axes and create a single legend
lines = line1 + line2 + line3
labels = [line.get_label() for line in lines]
ax1.legend(lines, labels, loc='lower left')
# Show the plot
plt.title("Training History")
plt.show()
输出:
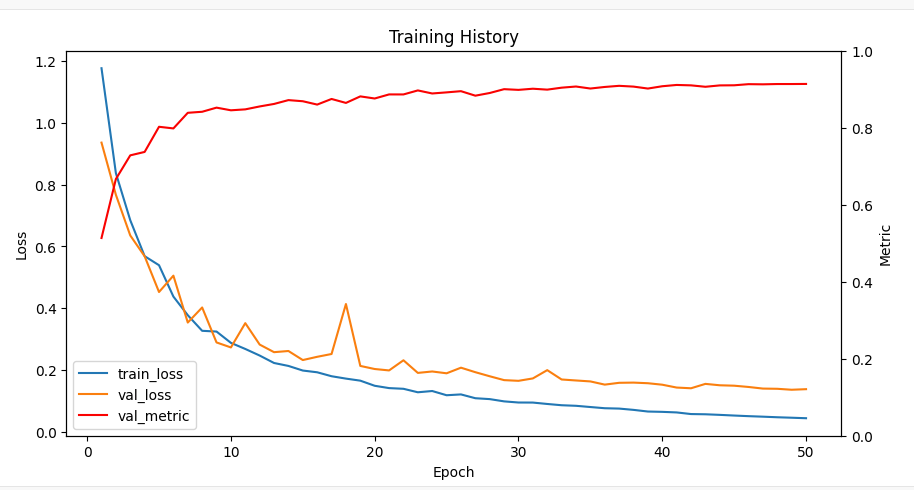
3.10模型保存和性能测试
def save_model(model,filepath):
# Saves all parameters that were changed during finetuning
# Create a dictionary to hold the non-frozen parameters
non_frozen_params = {}
# Iterate through all the model parameters
for param_name, param in model.named_parameters():
# If the parameter has requires_grad=True, add it to the dictionary
if param.requires_grad:
non_frozen_params[param_name] = param
# Save only the finetuned parameters
torch.save(non_frozen_params, filepath)
def load_model(checkpoint, filepath, num_labels=1, mixed = True, full = False, deepspeed = False):
# Creates a new PT5 model and loads the finetuned weights from a file
# load model
if"esm"in checkpoint:
model, tokenizer = load_esm_model(checkpoint, num_labels, mixed, full, deepspeed)
else:
model, tokenizer = load_T5_model(checkpoint, num_labels, mixed, full, deepspeed)
# Load the non-frozen parameters from the saved file
non_frozen_params = torch.load(filepath, weights_only=True)
# Assign the non-frozen parameters to the corresponding parameters of the model
for param_name, param in model.named_parameters():
if param_name in non_frozen_params:
param.data = non_frozen_params[param_name].data
return tokenizer, model
save_model(model, "./ESM2_8M_GB1_finetuned.pth")
tokenizer, model_reload = load_model(checkpoint, "./ESM2_8M_GB1_finetuned.pth", num_labels=1)
预测:
# For this we import the "three_vs_rest" GB1 dataset again
# from https://github.com/J-SNACKKB/FLIP
import requests
import zipfile
from io import BytesIO
# Download the zip file from GitHub
url = 'https://github.com/J-SNACKKB/FLIP/raw/main/splits/gb1/splits.zip'
response = requests.get(url)
zip_file = zipfile.ZipFile(BytesIO(response.content))
# Load the `three_vs_rest.csv` file into a pandas dataframe
with zip_file.open('splits/three_vs_rest.csv') as file:
df = pd.read_csv(file)
# Select only test data
my_test=df[df.set=="test"]
# Set column names to "sequence" and "label"
my_test.columns=["sequence","label"]+list(my_test.columns[2:])
# Drop unneeded columns
my_test=my_test[["sequence","label"]]
print(my_test.head(5))
# Preprocess sequences
my_test["sequence"]=my_test["sequence"].str.replace('|'.join(["O","B","U","Z"]),"X",regex=True)
my_test['sequence']=my_test.apply(lambda row : " ".join(row["sequence"]), axis = 1)
#Use reloaded model
model = model_reload
del model_reload
# Set the device to use
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
model.to(device)
# create Dataset
test_set=create_dataset(tokenizer,list(my_test['sequence']),list(my_test['label']))
# make compatible with torch DataLoader
test_set = test_set.with_format("torch", device=device)
# Create a dataloader for the test dataset
test_dataloader = DataLoader(test_set, batch_size=16, shuffle=False)
# Put the model in evaluation mode
model.eval()
# Make predictions on the test dataset
predictions = []
with torch.no_grad():
for batch in tqdm(test_dataloader):
input_ids = batch['input_ids'].to(device)
attention_mask = batch['attention_mask'].to(device)
#add batch results(logits) to predictions
predictions += model.float()(input_ids, attention_mask=attention_mask).logits.tolist()
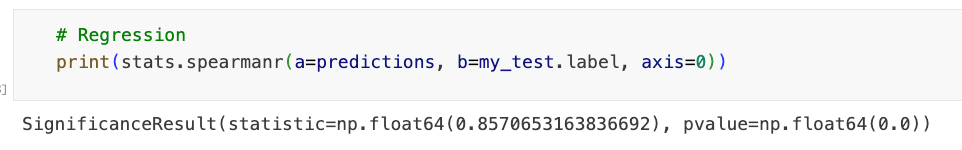
print(stats.spearmanr(a=predictions, b=my_test.label, axis=0))
输出:
-完-
本文参与 腾讯云自媒体同步曝光计划,分享自微信公众号。
原始发表:2026-03-23,如有侵权请联系 cloudcommunity@tencent.com 删除
评论
登录后参与评论
推荐阅读

