网络药理学—数据库获取疾病靶点并与中药靶点交集
原创网络药理学—数据库获取疾病靶点并与中药靶点交集
在中药复方的网络药理学分析中,比如说我们研究某某复方对某某疾病的有效果的原因。首先获取复方中中药成份中满足条件的小分子及其对应靶点,参考网络药理学—基于网络爬虫获取复方中药活性小分子及其对应靶点,第二步就是获取疾病靶点。然后对两方面靶点取交集,这些交集靶点就是目标靶点,后续的富集分析和PPI网络建立均以此为基础。
1 获取某疾病的医学主题词
首先在医学主题词网站https://meshb.nlm.nih.gov/中获取所研究疾病的最通用的名称,根据这个名称再去各个数据库中检索疾病靶点
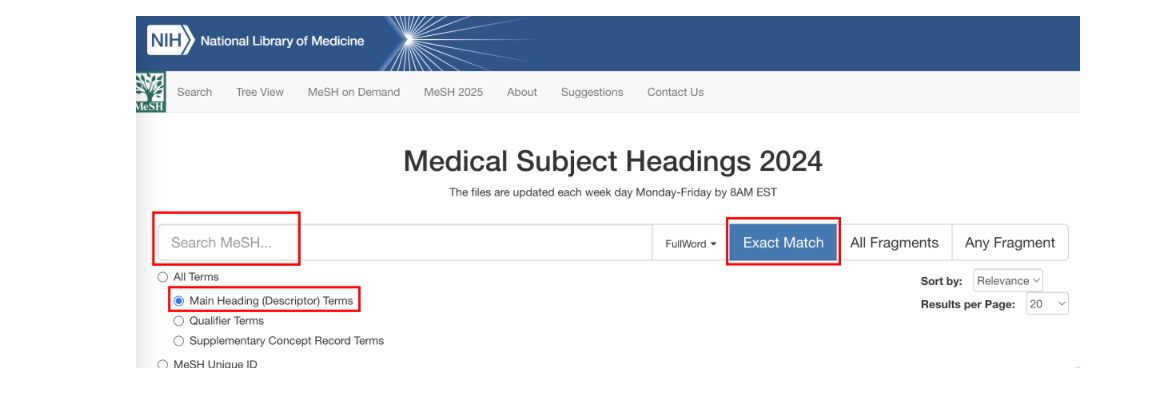
2 数据库获取疾病靶点
去各大数据库中下载疾病靶点,常见数据库如《滋肾清热利湿化瘀方治疗多囊卵巢综合征的网络药理学作用机制及实验研究》这篇文章中介绍的这些。一些疾病在部分数据库中检索不到属于正常,主要是OMIM、GeneCard、DisgeNET这三个数据库。
相关表格下载参考:https://www.bilibili.com/video/BV1yb421b7K6?spm_id_from=333.788.videopod.episodes&vd_source=7e83cb2510516bdff59ccf808d022aa0&p=3,后续我重写的代码,进行整理。

3 整理疾病靶点和获取中药疾病靶点交集
将每个疾病数据库中获取的疾病靶点取并集去重后,作为总体疾病靶点。再与第一步的target.csv文件中靶点取交集即可。执行下面这个脚本
3.Drug_case_gene.R
rm(list = ls())
options(stringsAsFactors = F)
library(readxl)
library(RColorBrewer)
library(data.table)
#### Drug target
{
gene_drug_target_df <- read.table("table/target.csv",sep=",",header=T)
gene_drug_target <- c()
for (genes in gene_drug_target_df$Gene.Names){
gene_drug_target <- c(gene_drug_target,strsplit(genes," ")[[1]])
}
gene_drug_target = unique(gene_drug_target)
}
#### Case target
{
##OMIM
gene_omim <- c()
gene_omim_df <- read_excel("table/case/OMIM-Gene-Map-Retrieval.xlsx")
for (genes in gene_omim_df$`Gene/Locus`){
gene_omim <- c(gene_omim,strsplit(genes,",")[[1]])
}
gene_omim = unique(gene_omim)
length(gene_omim)
gene_omim <- gene_omim[!is.na(gene_omim)]
length(gene_omim)#211
gene_genecard <- gene_genecard[!is.na(gene_genecard)]
gene_disgnet <- gene_disgnet[!is.na(gene_disgnet)]
##Genecard
gene_genecard <- c()
gene_genecard_df <- fread("table/case/GeneCards-SearchResults.csv",data.table = F)
dim(gene_genecard_df)#1527 8
#筛选阈值1或者10
gene_genecard_df <- gene_genecard_df[gene_genecard_df$`Relevance score` > 1,]
dim(gene_genecard_df)#1473 8
gene_genecard = unique(gene_genecard_df$`Gene Symbol`)
length(gene_genecard)
gene_genecard <- gene_genecard[!is.na(gene_genecard)]
length(gene_genecard)#1473
##Disgnet
gene_disgnet <- c()
gene_disgnet_df <- read_excel("table/case/DISEASES_Summary_GDA_CURATED_C3839507-C0085215-C0025322-C4552079.xlsx")
gene_disgnet = unique(gene_disgnet_df$gene)
length(gene_disgnet)
gene_disgnet <- gene_disgnet[!is.na(gene_disgnet)]
length(gene_disgnet)#141
## 数据库case target Venn
{
library(venn)
gene_sets <- list(
OMIM = gene_omim,
GeneCards = gene_genecard,
DisGeNET = gene_disgnet
)
mycol=c("#029149","#E0367A","#5D90BA","#431A3D","#91612D",
"#FFD121","#D8D155","#223D6C","#D20A13","#088247",
"#11AA4D","#7A142C","#5D90BA","#64495D","#7CC767")
pdf(file="figure/venn_case_target_gene.pdf",width=5,height=5)
venn(gene_sets,col=mycol[1:length(gene_sets)],zcolor=mycol[1:length(gene_sets)],box=F,ilabels = "counts")
#在 line 参数中,负值会将标题向上移动,正值会向下移动。可以调整 line 的值,直到标题完全显示。
title(main = "DOR Target Gene", cex.main = 1.5, font.main = 2, col.main = "black",line=-1)
dev.off()
}
gene_case_target <- unique(c(gene_omim,gene_genecard,gene_disgnet))
length(gene_case_target)#1744
}
#### Drug target 与 Case target 交集
{
Drug_case_target <- intersect(gene_drug_target,gene_case_target)
length(Drug_case_target)#128
#韦恩图
gene_sets <- list(
Drug_target = gene_drug_target,
Case_target = gene_case_target
)
pdf(file="figure/venn_Drug_case_target.pdf",width=5,height=5)
venn(gene_sets,col=mycol[1:length(gene_sets)],zcolor=mycol[1:length(gene_sets)],box=F,ilabels = "counts")
title(main = "DOR Target Gene", cex.main = 1.5, font.main = 2, col.main = "black",line=-1)
dev.off()
}
### 保存
write.csv(gene_drug_target,file = "../Table+Figure/T1.drug_target_gene.csv")
write.csv(gene_case_target,file = "../Table+Figure/T2.case_target_gene.csv")
write.csv(Drug_case_target,file = "../Table+Figure/T3.drug_case_target_intersect.csv")
save(gene_drug_target,gene_case_target,Drug_case_target,file = "Rdata/Drug_case_target.Rdata")得到两张图,第一张图疾病数据库获取的靶点交集情况
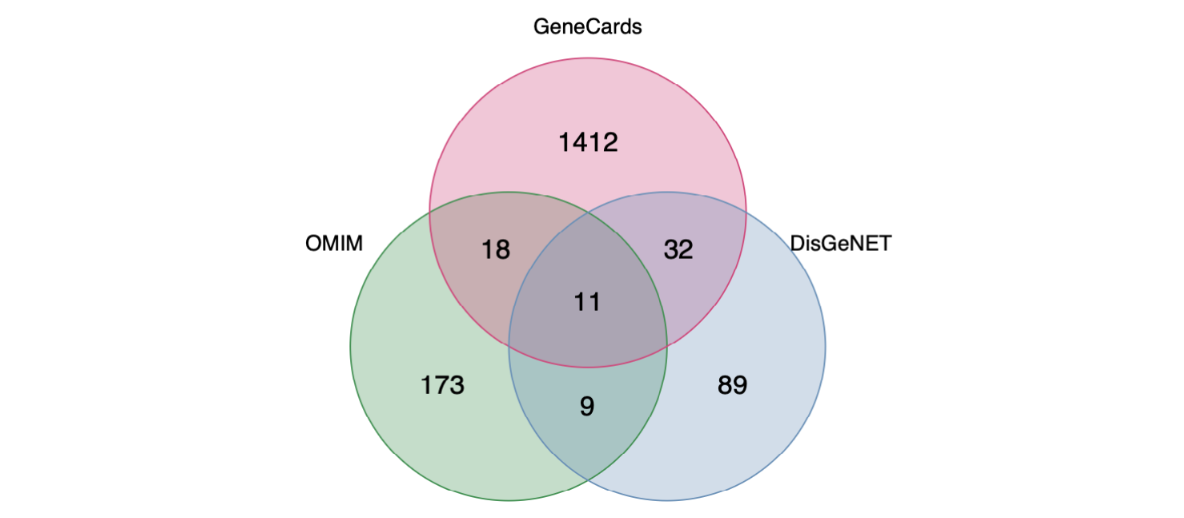
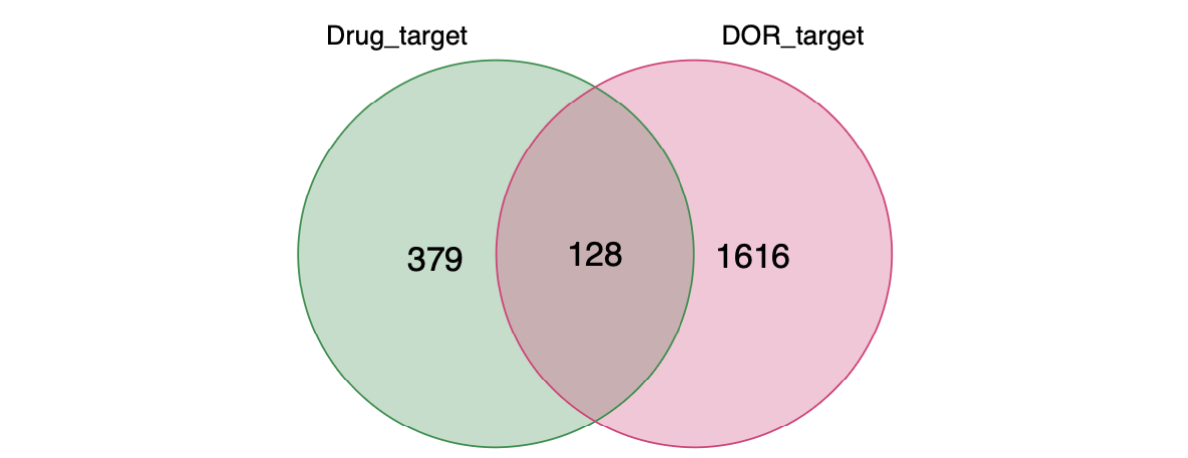
第二张图,疾病靶点并与中药靶点交集,即得到128个交集基因,后续分析全部基于这128个交集基因。
原创声明:本文系作者授权腾讯云开发者社区发表,未经许可,不得转载。
如有侵权,请联系 cloudcommunity@tencent.com 删除。
原创声明:本文系作者授权腾讯云开发者社区发表,未经许可,不得转载。
如有侵权,请联系 cloudcommunity@tencent.com 删除。
评论
登录后参与评论
推荐阅读
目录


