相关性热图、圈图、弦图(笔记)
下面是一年前的马拉松课程学员的分享
生信技能树优秀学员笔记,代码主要来自生信技能树的直播课程,加上自己的理解和发挥。
数据准备:
如下所示,可以看到有多个样品,每个样品都有多个基因表达量,这个时候我们比较关心的是这些基因的表达量相关性(在多个样品),基因与基因之间有两两组合相关性:
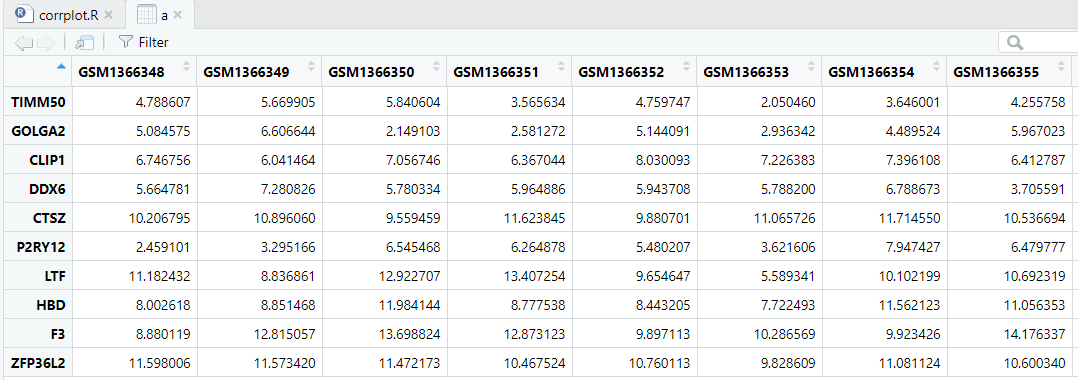
M: 很容易计算基因之间的相关性矩阵
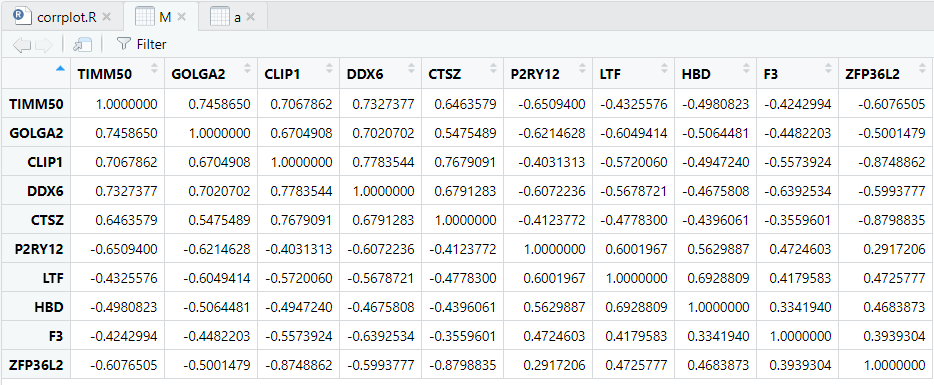
#感兴趣基因/样本的相关性图----
rm(list = ls())
load(file = "step1output.Rdata")
load(file = "step4output.Rdata")
load(file = 'step2output.Rdata')
library(dplyr)
library(ggplot2)
library(pheatmap)
library(corrplot)
dat = deg[!duplicated(deg$symbol),]
exp = exp[dat$probe_id,]
rownames(exp) = dat$symbol
g = c("TIMM50", "GOLGA2", "CLIP1", "DDX6", "CTSZ", "P2RY12", "LTF",
"HBD", "F3", "ZFP36L2")
a = exp[g,]
M = cor(t(exp[g,])) #cor()函数可以对矩阵计算相关性(谁在列的位置就计算谁的相关性)
#画基因之间的相关性,cor函数后面的矩阵exp[g,]要以基因为列名(转置一下)
#画样本之间的相关性,cor函数后面的矩阵exp[g,]要以样本为列名(不要转置)
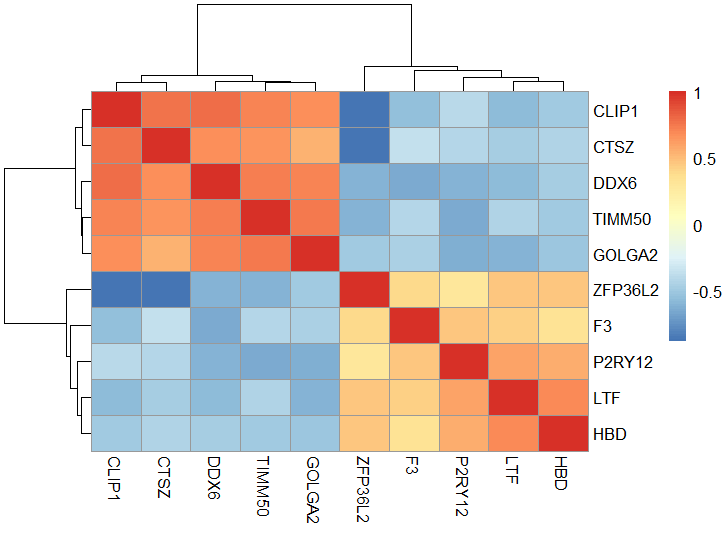
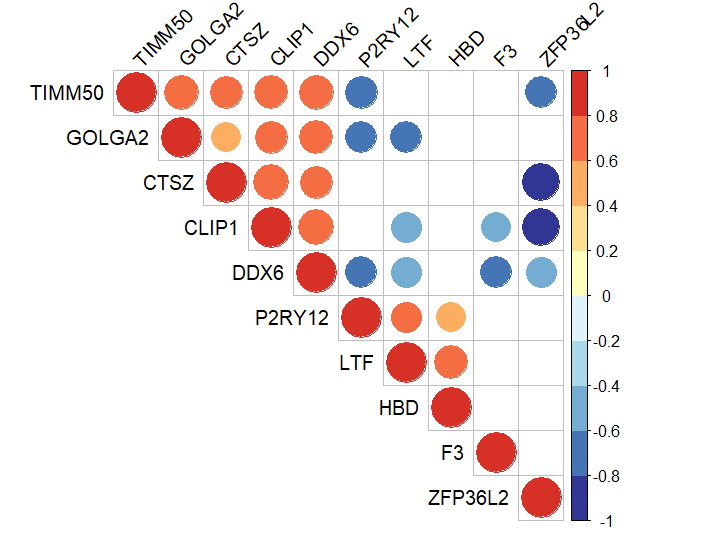
#相关性热图
pheatmap(M)
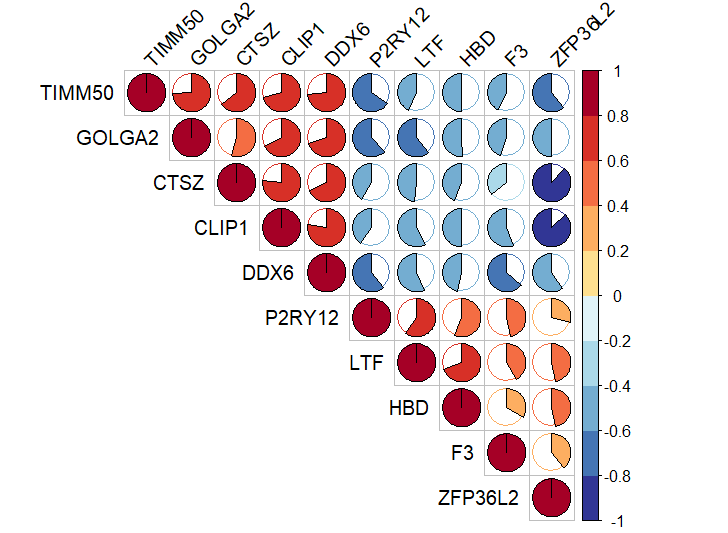
#相关性圆圈图
library(paletteer)
my_color = rev(paletteer_d("RColorBrewer::RdYlBu"))
my_color
my_color = colorRampPalette(my_color)(10) #整成10个色阶
# library(RColorBrewer)
# my_color=rev(brewer.pal(10, "RdYlBu"))
corrplot(M, type="upper",
method="pie",
order="hclust",
col=my_color,
tl.col="black",
#tl.pos = "d",
tl.srt=45
)
#corrplot不能赋值,和其他R包拼图就比较费劲
library(cowplot) #比patchwork包更强的拼图包
cor_plot <- recordPlot() #cowplot包里的recordPlot函数可以把这个相关性图抠下来强行赋值
# 拼图(相关性图属于另外一个拼图体系)
#load("pca_plot.Rdata")
pdf("cor_plot.pdf", width = 10, height = 10)
plot_grid(cor_plot)
#plot_grid(pca_plot,cor_plot) #cowplot包里的plot_grid()函数允许跨绘图体系拼图
dev.off()
## 圆圈视图
M = cor(t(a))
p.mat <- cor.mtest(t(a))$p
library(paletteer)
my_color = rev(paletteer_d("RColorBrewer::RdYlBu")[-1])
my_color = colorRampPalette(my_color)(10)
png("plot/corrplot.png",height = 1000,width = 1000)
corrplot(M, type="upper",
order="hclust",
col = my_color,
p.mat = p.mat,
sig.level = 0.01,
insig = "blank",
tl.col = "black",
tl.srt=45)
dev.off()
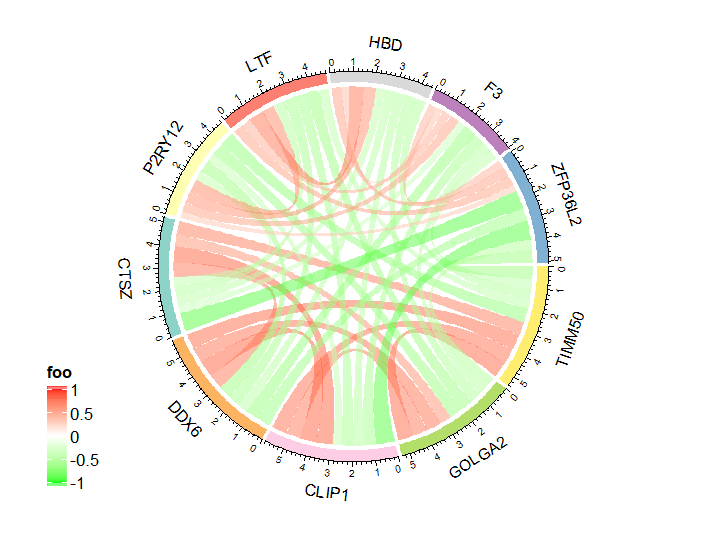
相关性弦图:连线就代表了基因间表达值的相关性信息,红色代表正相关,绿色代表负相关,颜色越深或连线越粗代表相关强度越高。 一行是一个连接关系,第一列和第二列是要连接的两边,第三列是连接的强度。
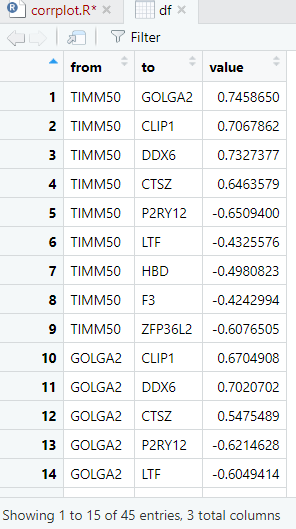
## 相关性弦图
library(circlize)
library(tidyr)
library(tibble)
library(ComplexHeatmap)
mat = M
df = mat %>%
as.data.frame() %>%
rownames_to_column(var = "from") %>%
pivot_longer(names_to = "to",
values_to = "value",
cols = 2:(nrow(mat)+1) #有10个基因,这里就是11
)
df = df[df$from != df$to,]
df = df[!duplicated(df$value),]
#自定义边的颜色
library(RColorBrewer)
col_sample = c(brewer.pal(n = 12,name = "Set3")
#brewer.pal(n = 8,name = "Set2")
)
col_sample
border_color <- sample(col_sample,nrow(mat))
#border_color = c("#66C2A5", "#FC8D62", "#8DA0CB", "#E78AC3")
#根据相关性大小展示连线的颜色范围:红 白 绿
col = colorRamp2(c(-1, 0, 1), c('green', 'white', 'red'), transparency = 0.5)
# range(mat)
# col <- colorRamp2(range(mat), c("#FA0A0A", "#FFFFFF"))
# ComplexHeatmapt添加图例
lgd = Legend(col_fun = col, title = "foo")
ComplexHeatmap:::width(lgd)
ComplexHeatmap:::height(lgd)
pushViewport(viewport(width = 0.9, height = 0.9))
grid.rect() # border
draw(lgd, x = unit(1, "cm"), y = unit(1, "cm"), just = c("left", "bottom"))
# draw(lgd, x = unit(0.5, "npc"), y = unit(0.5, "npc"))
# draw(lgd, x = unit(1, "npc"), y = unit(1, "npc"), just = c("right", "top"))
popViewport()
chordDiagram(x = df,
directional = 0, #表示线条的方向,0代表没有方向,1代表正向,-1代表反向,2代表双向
grid.col = border_color, #圈的颜色
col = col, #弦的颜色
annotationTrack = c('grid', 'name', 'axis'), #绘制外周圆弧区,显示名称和刻度轴
annotationTrackHeight = c(0.05, 0.1),#外面一圈的宽度
transparency = 0.25#线条的透明度
)
以下是小洁老师看到我的需求后发来的资料投喂: 弦图的边距问题: R circlize - 绘制边距和绘制区域- circlize - 码客 (oomake.com) 弦图函数的书: Chapter 16 A complex example of Chord diagram | Circular Visualization in R (jokergoo.github.io) 弦图图例用complexheatmap实现: Chapter 5 Legends | ComplexHeatmap Complete Reference (jokergoo.github.io
本文参与 腾讯云自媒体同步曝光计划,分享自微信公众号。
原始发表:2023-02-19,如有侵权请联系 cloudcommunity@tencent.com 删除
评论
登录后参与评论
推荐阅读

